Probe CUST_28360_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28360_PI426222305 | JHI_St_60k_v1 | DMT400044438 | GGCTAGTATTCTTTGTTTTTCTATACCATATACGACTACGATCAATAAGAGGCACTGATA |
All Microarray Probes Designed to Gene DMG400017246
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28270_PI426222305 | JHI_St_60k_v1 | DMT400044437 | TTGTTGTGAACATCGGAGATCTTCTGCAGCTTATATCAAATGACAAGTACATAAGTGTTG |
| CUST_28360_PI426222305 | JHI_St_60k_v1 | DMT400044438 | GGCTAGTATTCTTTGTTTTTCTATACCATATACGACTACGATCAATAAGAGGCACTGATA |
Microarray Signals from CUST_28360_PI426222305
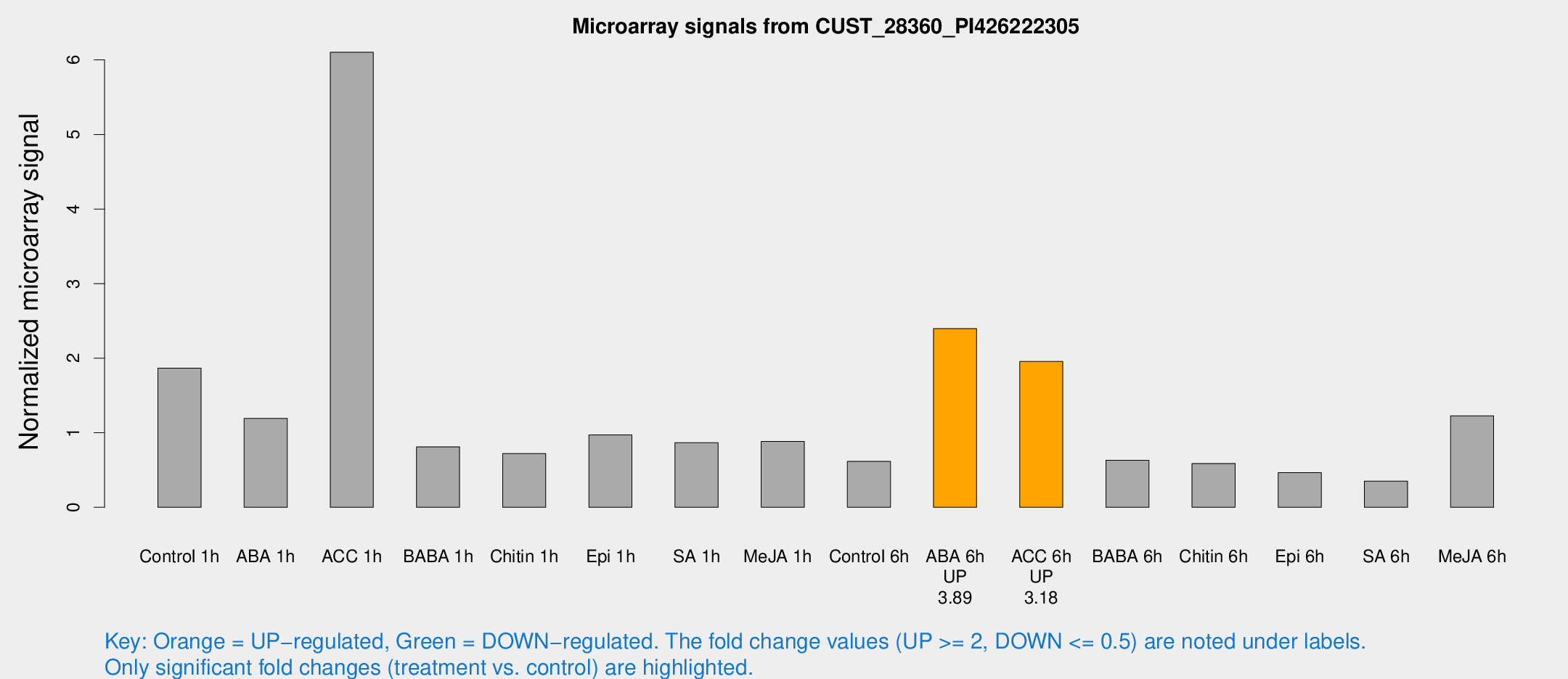
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 45.953 | 6.21783 | 1.867 | 0.262458 |
| ABA 1h | 28.1878 | 7.73135 | 1.19171 | 0.361031 |
| ACC 1h | 163.884 | 43.9939 | 6.10317 | 1.42759 |
| BABA 1h | 24.4065 | 9.51359 | 0.810035 | 0.452875 |
| Chitin 1h | 17.1658 | 5.25862 | 0.720892 | 0.200624 |
| Epi 1h | 20.7171 | 3.56896 | 0.972373 | 0.177135 |
| SA 1h | 25.0743 | 9.8693 | 0.866562 | 0.356796 |
| Me-JA 1h | 18.0541 | 3.70325 | 0.884763 | 0.195045 |
| Control 6h | 16.1585 | 4.8444 | 0.615505 | 0.177938 |
| ABA 6h | 63.3783 | 10.3335 | 2.39569 | 0.344697 |
| ACC 6h | 54.8329 | 6.25875 | 1.95591 | 0.188992 |
| BABA 6h | 19.4072 | 7.3604 | 0.629866 | 0.216487 |
| Chitin 6h | 17.4229 | 5.59805 | 0.5881 | 0.294793 |
| Epi 6h | 13.3454 | 4.22249 | 0.465137 | 0.173286 |
| SA 6h | 8.58353 | 3.71796 | 0.351192 | 0.16053 |
| Me-JA 6h | 34.8464 | 11.9953 | 1.22628 | 0.489465 |
Source Transcript PGSC0003DMT400044438 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G06620.1 | +1 | 3e-120 | 336 | 167/273 (61%) | 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein | chr1:2025618-2027094 FORWARD LENGTH=365 |
