Probe CUST_28270_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28270_PI426222305 | JHI_St_60k_v1 | DMT400044437 | TTGTTGTGAACATCGGAGATCTTCTGCAGCTTATATCAAATGACAAGTACATAAGTGTTG |
All Microarray Probes Designed to Gene DMG400017246
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28270_PI426222305 | JHI_St_60k_v1 | DMT400044437 | TTGTTGTGAACATCGGAGATCTTCTGCAGCTTATATCAAATGACAAGTACATAAGTGTTG |
| CUST_28360_PI426222305 | JHI_St_60k_v1 | DMT400044438 | GGCTAGTATTCTTTGTTTTTCTATACCATATACGACTACGATCAATAAGAGGCACTGATA |
Microarray Signals from CUST_28270_PI426222305
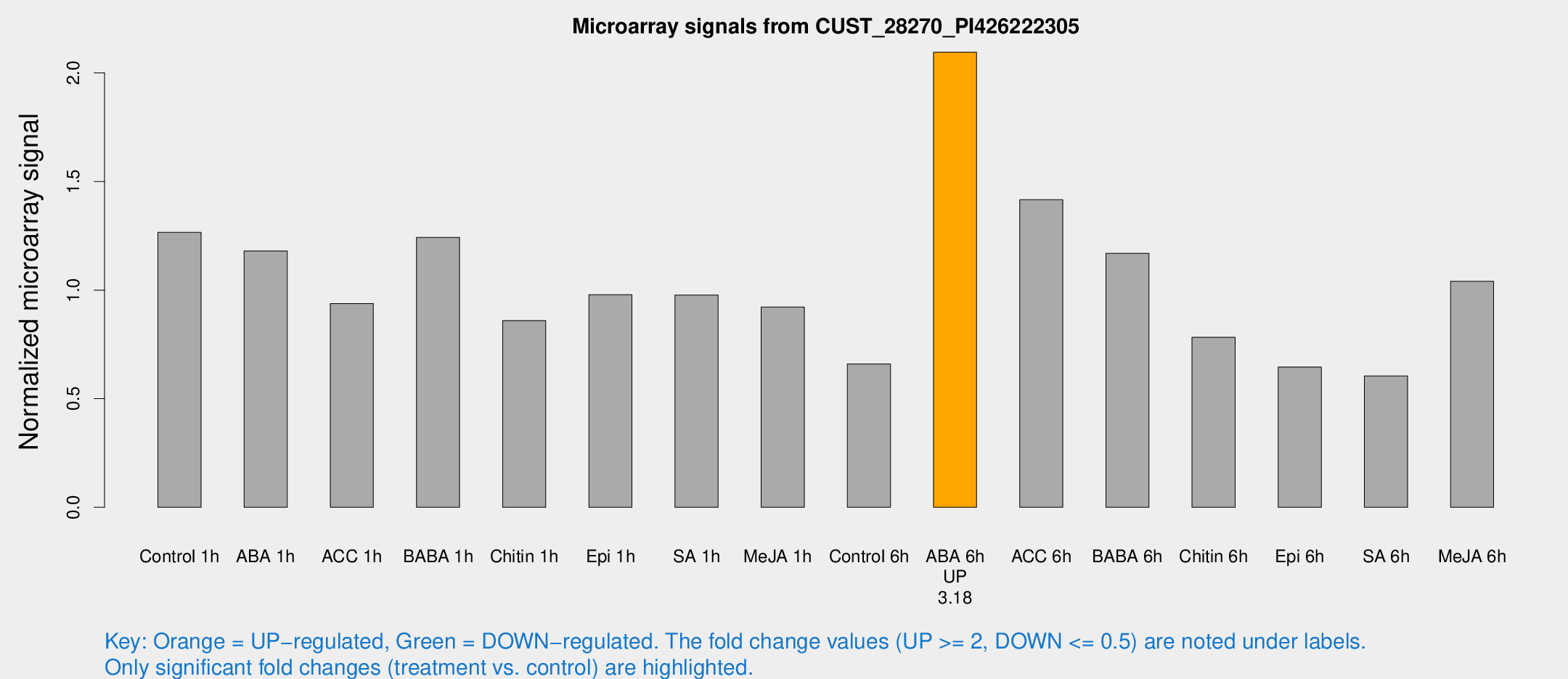
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1563.08 | 196.07 | 1.26592 | 0.111731 |
| ABA 1h | 1366.2 | 332.826 | 1.17957 | 0.295367 |
| ACC 1h | 2960.21 | 1484.45 | 0.937554 | 5.33626 |
| BABA 1h | 1614.05 | 481.507 | 1.24253 | 0.304816 |
| Chitin 1h | 996.338 | 252.049 | 0.859662 | 0.147127 |
| Epi 1h | 1057.72 | 164.788 | 0.979055 | 0.156593 |
| SA 1h | 1279.13 | 252.285 | 0.97674 | 0.171506 |
| Me-JA 1h | 944.562 | 156.495 | 0.921611 | 0.117114 |
| Control 6h | 874.798 | 232.385 | 0.659724 | 0.15008 |
| ABA 6h | 2805.99 | 485.521 | 2.09497 | 0.320565 |
| ACC 6h | 2020.82 | 315.61 | 1.41588 | 0.0818097 |
| BABA 6h | 1629.81 | 254.506 | 1.16939 | 0.16074 |
| Chitin 6h | 1020.31 | 81.9808 | 0.782675 | 0.0903383 |
| Epi 6h | 895.645 | 92.4934 | 0.645783 | 0.0374227 |
| SA 6h | 755.888 | 143.808 | 0.604413 | 0.0598875 |
| Me-JA 6h | 1282 | 167.229 | 1.04045 | 0.0601615 |
Source Transcript PGSC0003DMT400044437 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G06620.1 | +1 | 2e-153 | 444 | 219/356 (62%) | 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein | chr1:2025618-2027094 FORWARD LENGTH=365 |
