Probe CUST_28145_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28145_PI426222305 | JHI_St_60k_v1 | DMT400004535 | CATTCGTCGCCCTGGGTAAAGCATCAAATTATTTCTGCAATTCAAACCTATAATTGATAA |
All Microarray Probes Designed to Gene DMG400001803
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28162_PI426222305 | JHI_St_60k_v1 | DMT400004533 | CAATTAGATATGATTGATGGATGAAAAGATCCATAGAAAAGAAGCTCACGGTCATGATTC |
| CUST_28145_PI426222305 | JHI_St_60k_v1 | DMT400004535 | CATTCGTCGCCCTGGGTAAAGCATCAAATTATTTCTGCAATTCAAACCTATAATTGATAA |
| CUST_28065_PI426222305 | JHI_St_60k_v1 | DMT400004534 | CAATTAGATATGATTGATGGATGAAAAGATCCATAGAAAAGAAGCTCACGGTCATGATTC |
Microarray Signals from CUST_28145_PI426222305
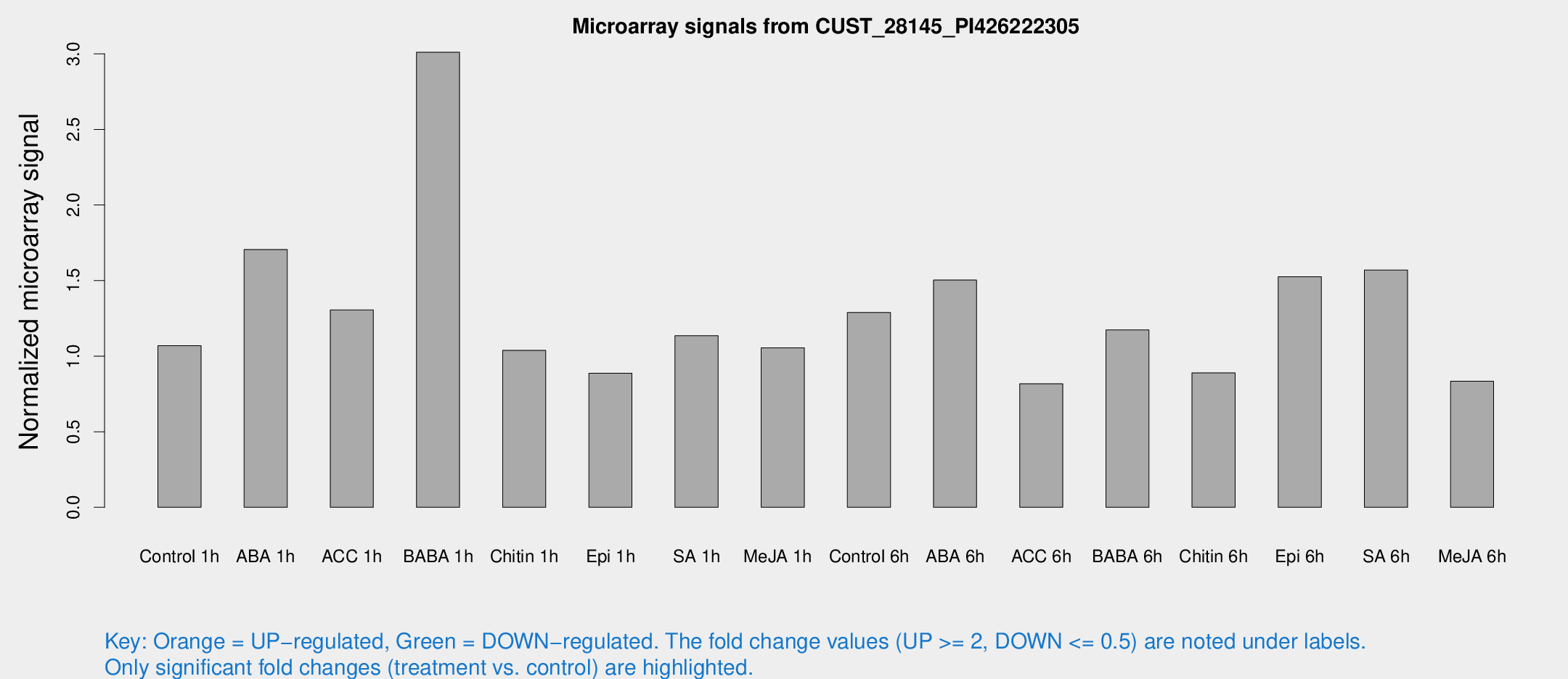
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 8.96635 | 3.55619 | 1.06951 | 0.510155 |
| ABA 1h | 13.4076 | 5.13062 | 1.70537 | 0.633376 |
| ACC 1h | 10.807 | 4.55561 | 1.30554 | 0.617855 |
| BABA 1h | 23.7243 | 6.30281 | 3.01062 | 1.25828 |
| Chitin 1h | 7.55744 | 3.53425 | 1.03827 | 0.525793 |
| Epi 1h | 5.83869 | 3.40071 | 0.887097 | 0.514616 |
| SA 1h | 9.60205 | 3.55722 | 1.13517 | 0.501139 |
| Me-JA 1h | 6.56333 | 3.63051 | 1.05476 | 0.579428 |
| Control 6h | 11.8033 | 5.34877 | 1.289 | 0.580508 |
| ABA 6h | 12.5561 | 3.85902 | 1.50353 | 0.495421 |
| ACC 6h | 7.29208 | 4.30554 | 0.817325 | 0.470923 |
| BABA 6h | 10.1052 | 4.10425 | 1.17354 | 0.501875 |
| Chitin 6h | 7.17421 | 4.06264 | 0.889217 | 0.502538 |
| Epi 6h | 13.3969 | 4.34571 | 1.52474 | 0.551438 |
| SA 6h | 13.1502 | 4.42896 | 1.56953 | 0.817559 |
| Me-JA 6h | 6.29324 | 3.64959 | 0.834177 | 0.483544 |
Source Transcript PGSC0003DMT400004535 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | None | - | - | - | - | - |
