Probe CUST_2803_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_2803_PI426222305 | JHI_St_60k_v1 | DMT400033249 | GATGGTGATGGATTTGTTAATTTTGAAGAGTTTAAACAAATGATGGCTGCTGGATGCAAT |
All Microarray Probes Designed to Gene DMG400012770
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_2897_PI426222305 | JHI_St_60k_v1 | DMT400033250 | GTAGGATGTCAATGTTATGAAAGCAAGTAGGTAACTCCATATTTAGCATAATTCTCCTCT |
| CUST_2803_PI426222305 | JHI_St_60k_v1 | DMT400033249 | GATGGTGATGGATTTGTTAATTTTGAAGAGTTTAAACAAATGATGGCTGCTGGATGCAAT |
Microarray Signals from CUST_2803_PI426222305
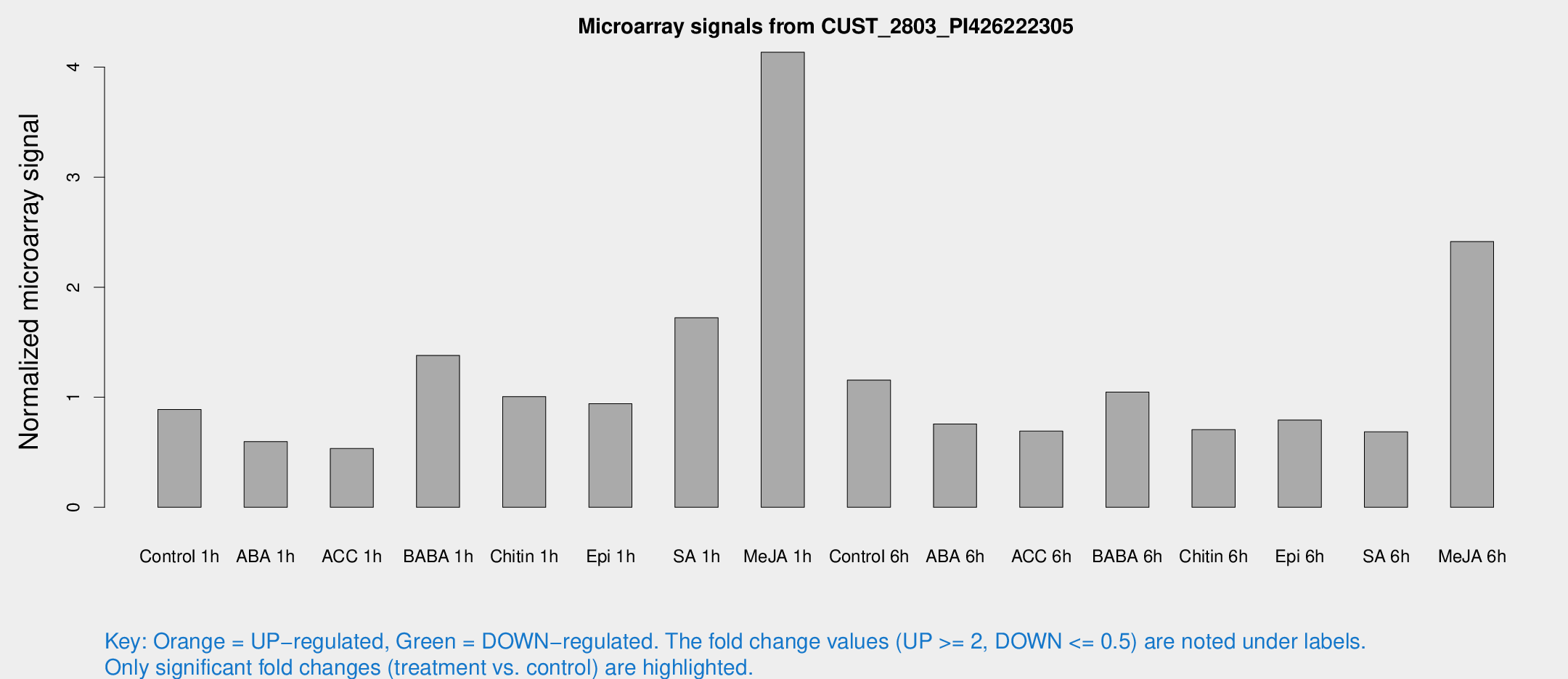
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 230.142 | 52.4776 | 0.889211 | 0.145729 |
| ABA 1h | 155.524 | 59.9133 | 0.597076 | 0.252912 |
| ACC 1h | 152.559 | 45.5669 | 0.534147 | 0.173182 |
| BABA 1h | 352.557 | 87.2764 | 1.38023 | 0.261706 |
| Chitin 1h | 226.811 | 32.0269 | 1.00621 | 0.119815 |
| Epi 1h | 201.537 | 16.0419 | 0.941272 | 0.0711074 |
| SA 1h | 448.396 | 73.273 | 1.72205 | 0.231363 |
| Me-JA 1h | 901.008 | 262.366 | 4.13468 | 0.848049 |
| Control 6h | 319.89 | 97.9759 | 1.15572 | 0.322893 |
| ABA 6h | 207.989 | 46.7744 | 0.757568 | 0.128204 |
| ACC 6h | 195.654 | 12.1703 | 0.691845 | 0.126244 |
| BABA 6h | 294.403 | 44.094 | 1.04684 | 0.11605 |
| Chitin 6h | 185.993 | 16.0275 | 0.705371 | 0.0858505 |
| Epi 6h | 227.256 | 39.3111 | 0.792616 | 0.206993 |
| SA 6h | 249.238 | 106.679 | 0.685538 | 0.609746 |
| Me-JA 6h | 613.222 | 122.722 | 2.41434 | 0.287614 |
Source Transcript PGSC0003DMT400033249 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G24620.1 | +1 | 5e-36 | 125 | 71/146 (49%) | EF hand calcium-binding protein family | chr1:8723893-8724453 REVERSE LENGTH=186 |
