Probe CUST_2897_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_2897_PI426222305 | JHI_St_60k_v1 | DMT400033250 | GTAGGATGTCAATGTTATGAAAGCAAGTAGGTAACTCCATATTTAGCATAATTCTCCTCT |
All Microarray Probes Designed to Gene DMG400012770
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_2897_PI426222305 | JHI_St_60k_v1 | DMT400033250 | GTAGGATGTCAATGTTATGAAAGCAAGTAGGTAACTCCATATTTAGCATAATTCTCCTCT |
| CUST_2803_PI426222305 | JHI_St_60k_v1 | DMT400033249 | GATGGTGATGGATTTGTTAATTTTGAAGAGTTTAAACAAATGATGGCTGCTGGATGCAAT |
Microarray Signals from CUST_2897_PI426222305
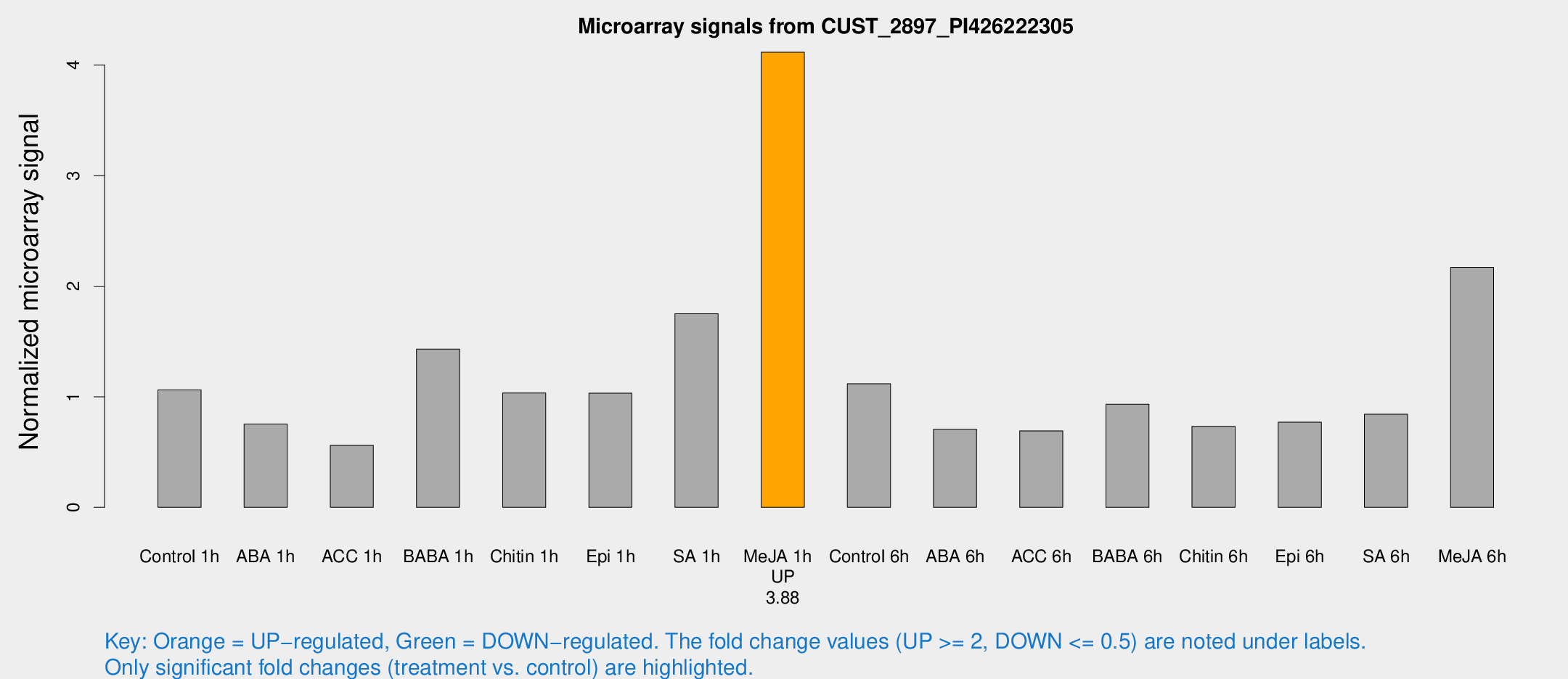
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 414.754 | 69.7043 | 1.06175 | 0.128357 |
| ABA 1h | 272.849 | 69.2737 | 0.753773 | 0.164404 |
| ACC 1h | 254.522 | 85.1211 | 0.56048 | 0.210443 |
| BABA 1h | 549.714 | 111.492 | 1.43081 | 0.178176 |
| Chitin 1h | 356.732 | 31.0199 | 1.03472 | 0.106141 |
| Epi 1h | 340.446 | 20.0239 | 1.03154 | 0.0605618 |
| SA 1h | 694.119 | 81.5166 | 1.75039 | 0.158065 |
| Me-JA 1h | 1324.57 | 257.175 | 4.1161 | 0.428237 |
| Control 6h | 456.196 | 112.732 | 1.11779 | 0.200606 |
| ABA 6h | 291.001 | 43.057 | 0.70507 | 0.0654992 |
| ACC 6h | 309.567 | 51.1022 | 0.691685 | 0.198411 |
| BABA 6h | 398.933 | 30.5924 | 0.931614 | 0.054723 |
| Chitin 6h | 296.391 | 17.6933 | 0.7313 | 0.0436443 |
| Epi 6h | 338.622 | 51.0815 | 0.770102 | 0.173191 |
| SA 6h | 384.174 | 134.456 | 0.841284 | 0.344113 |
| Me-JA 6h | 831.774 | 91.549 | 2.17078 | 0.125752 |
Source Transcript PGSC0003DMT400033250 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G24620.1 | +2 | 2e-33 | 125 | 71/146 (49%) | EF hand calcium-binding protein family | chr1:8723893-8724453 REVERSE LENGTH=186 |
