Probe CUST_2800_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_2800_PI426222305 | JHI_St_60k_v1 | DMT400000383 | AGTGCAGGTATCACTAAAACTCTTGAGAGGTACCAACGTTGTTGTCTTAATCCTCAAGAC |
All Microarray Probes Designed to Gene DMG400000136
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_2934_PI426222305 | JHI_St_60k_v1 | DMT400000382 | TTTTGCAGTATCACTAAAACTCTTGAGAGGTACCAACGTTGTTGTCTTAATCCTCAAGAC |
| CUST_2800_PI426222305 | JHI_St_60k_v1 | DMT400000383 | AGTGCAGGTATCACTAAAACTCTTGAGAGGTACCAACGTTGTTGTCTTAATCCTCAAGAC |
Microarray Signals from CUST_2800_PI426222305
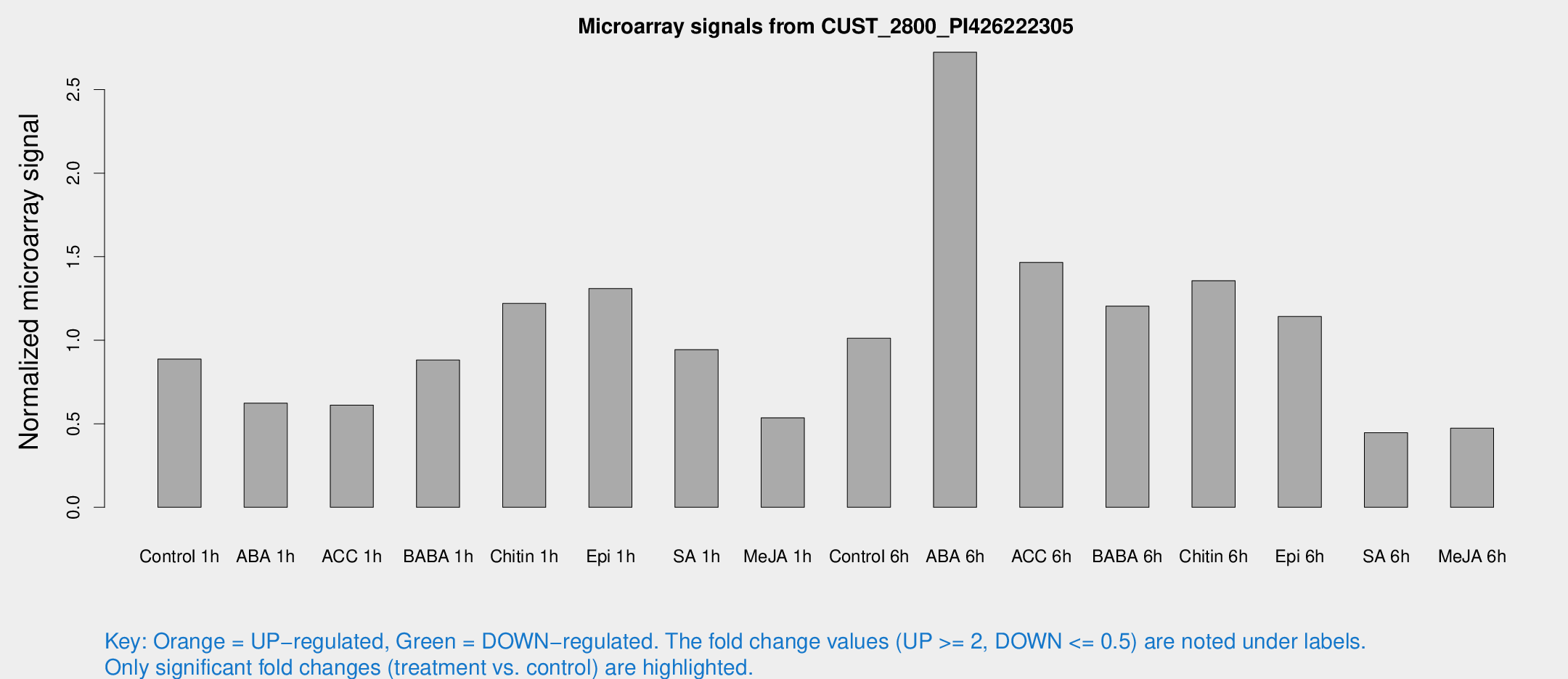
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 57.4781 | 10.7772 | 0.887318 | 0.132502 |
| ABA 1h | 34.6538 | 3.78171 | 0.623703 | 0.0773435 |
| ACC 1h | 72.915 | 41.4273 | 0.611074 | 1.00659 |
| BABA 1h | 56.6799 | 14.8726 | 0.88136 | 0.177065 |
| Chitin 1h | 71.4669 | 13.7077 | 1.2203 | 0.230844 |
| Epi 1h | 71.746 | 7.27015 | 1.30926 | 0.115006 |
| SA 1h | 61.3699 | 6.35529 | 0.943527 | 0.0952533 |
| Me-JA 1h | 28.5961 | 6.14323 | 0.535629 | 0.165208 |
| Control 6h | 98.0746 | 44.1379 | 1.01174 | 1.05244 |
| ABA 6h | 194.08 | 48.1567 | 2.72371 | 0.706866 |
| ACC 6h | 121.27 | 46.9383 | 1.46575 | 0.381008 |
| BABA 6h | 91.529 | 23.8544 | 1.20393 | 0.336437 |
| Chitin 6h | 91.389 | 9.35344 | 1.35627 | 0.164482 |
| Epi 6h | 82.61 | 12.9216 | 1.14266 | 0.146939 |
| SA 6h | 31.0066 | 9.79126 | 0.446208 | 0.238837 |
| Me-JA 6h | 30.9364 | 6.96089 | 0.473661 | 0.102386 |
Source Transcript PGSC0003DMT400000383 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G45650.1 | +1 | 9e-90 | 275 | 157/261 (60%) | AGAMOUS-like 6 | chr2:18804453-18806291 FORWARD LENGTH=252 |
