Probe CUST_2934_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_2934_PI426222305 | JHI_St_60k_v1 | DMT400000382 | TTTTGCAGTATCACTAAAACTCTTGAGAGGTACCAACGTTGTTGTCTTAATCCTCAAGAC |
All Microarray Probes Designed to Gene DMG400000136
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_2934_PI426222305 | JHI_St_60k_v1 | DMT400000382 | TTTTGCAGTATCACTAAAACTCTTGAGAGGTACCAACGTTGTTGTCTTAATCCTCAAGAC |
| CUST_2800_PI426222305 | JHI_St_60k_v1 | DMT400000383 | AGTGCAGGTATCACTAAAACTCTTGAGAGGTACCAACGTTGTTGTCTTAATCCTCAAGAC |
Microarray Signals from CUST_2934_PI426222305
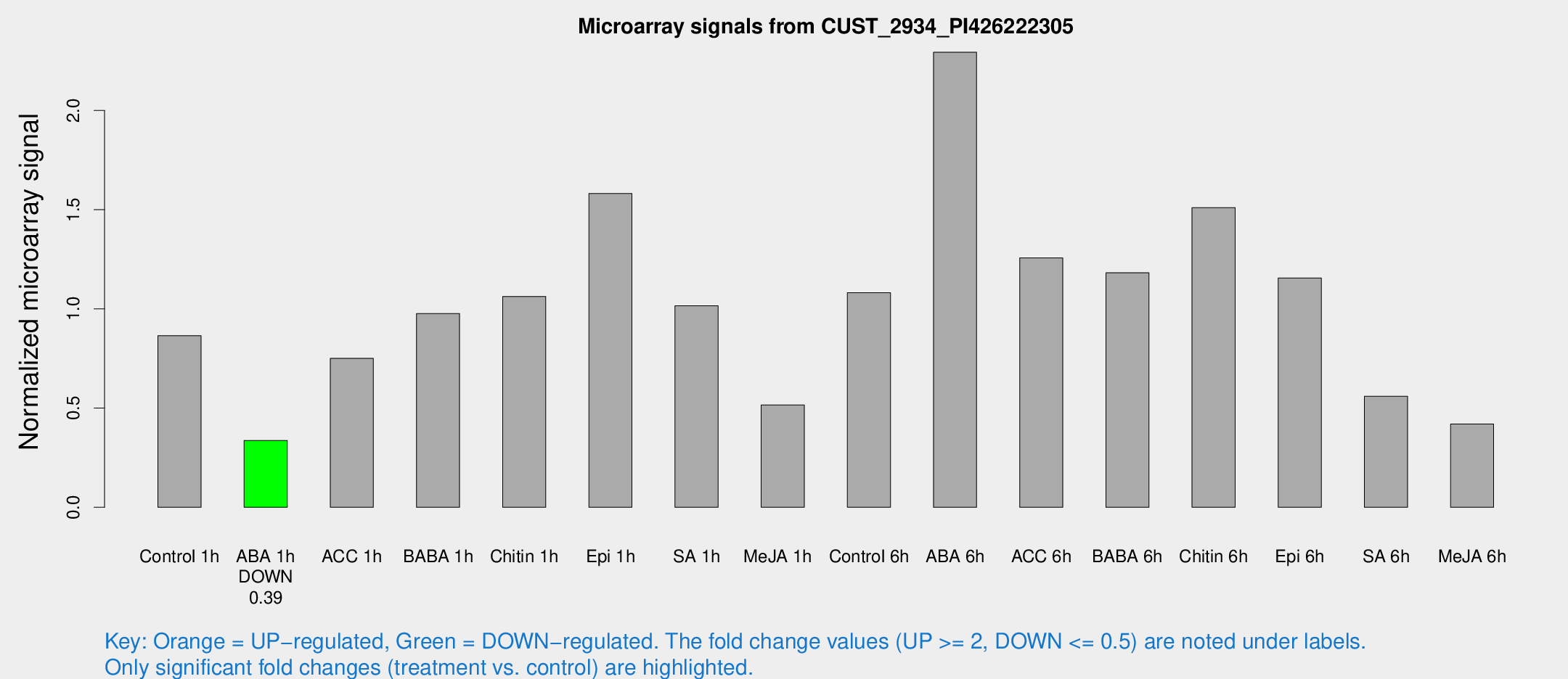
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 41.6054 | 4.87305 | 0.865196 | 0.112232 |
| ABA 1h | 14.4274 | 3.1722 | 0.337092 | 0.0779083 |
| ACC 1h | 50.303 | 23.0808 | 0.750826 | 0.527605 |
| BABA 1h | 47.7689 | 13.1541 | 0.976585 | 0.217311 |
| Chitin 1h | 48.5352 | 13.1891 | 1.0626 | 0.233688 |
| Epi 1h | 66.8097 | 10.8583 | 1.58125 | 0.256017 |
| SA 1h | 50.7647 | 8.60485 | 1.01572 | 0.226425 |
| Me-JA 1h | 23.7382 | 9.23295 | 0.516152 | 0.331334 |
| Control 6h | 85.6853 | 40.5305 | 1.08147 | 1.41537 |
| ABA 6h | 127.07 | 36.831 | 2.29353 | 0.732987 |
| ACC 6h | 82.7438 | 33.9598 | 1.25716 | 0.500916 |
| BABA 6h | 68.4413 | 19.9176 | 1.18198 | 0.343049 |
| Chitin 6h | 77.0857 | 8.35222 | 1.5106 | 0.143635 |
| Epi 6h | 63.0786 | 8.92817 | 1.15574 | 0.17947 |
| SA 6h | 27.3932 | 5.22688 | 0.559471 | 0.179434 |
| Me-JA 6h | 21.946 | 6.06417 | 0.419961 | 0.120643 |
Source Transcript PGSC0003DMT400000382 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G45650.1 | +2 | 4e-48 | 165 | 102/208 (49%) | AGAMOUS-like 6 | chr2:18804453-18806291 FORWARD LENGTH=252 |
