Probe CUST_27917_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_27917_PI426222305 | JHI_St_60k_v1 | DMT400082027 | TAACTTTAGGCATGGACAAATAATAGTTTGACATCTGCTGAACACAGAGTAGTACCAACA |
All Microarray Probes Designed to Gene DMG400032208
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_27917_PI426222305 | JHI_St_60k_v1 | DMT400082027 | TAACTTTAGGCATGGACAAATAATAGTTTGACATCTGCTGAACACAGAGTAGTACCAACA |
| CUST_28016_PI426222305 | JHI_St_60k_v1 | DMT400082026 | AACATCCCCTTGAGGTGCTGCCCTTTCTCTGTGAAAGAAATCACTAATGCTTTTTATGTG |
Microarray Signals from CUST_27917_PI426222305
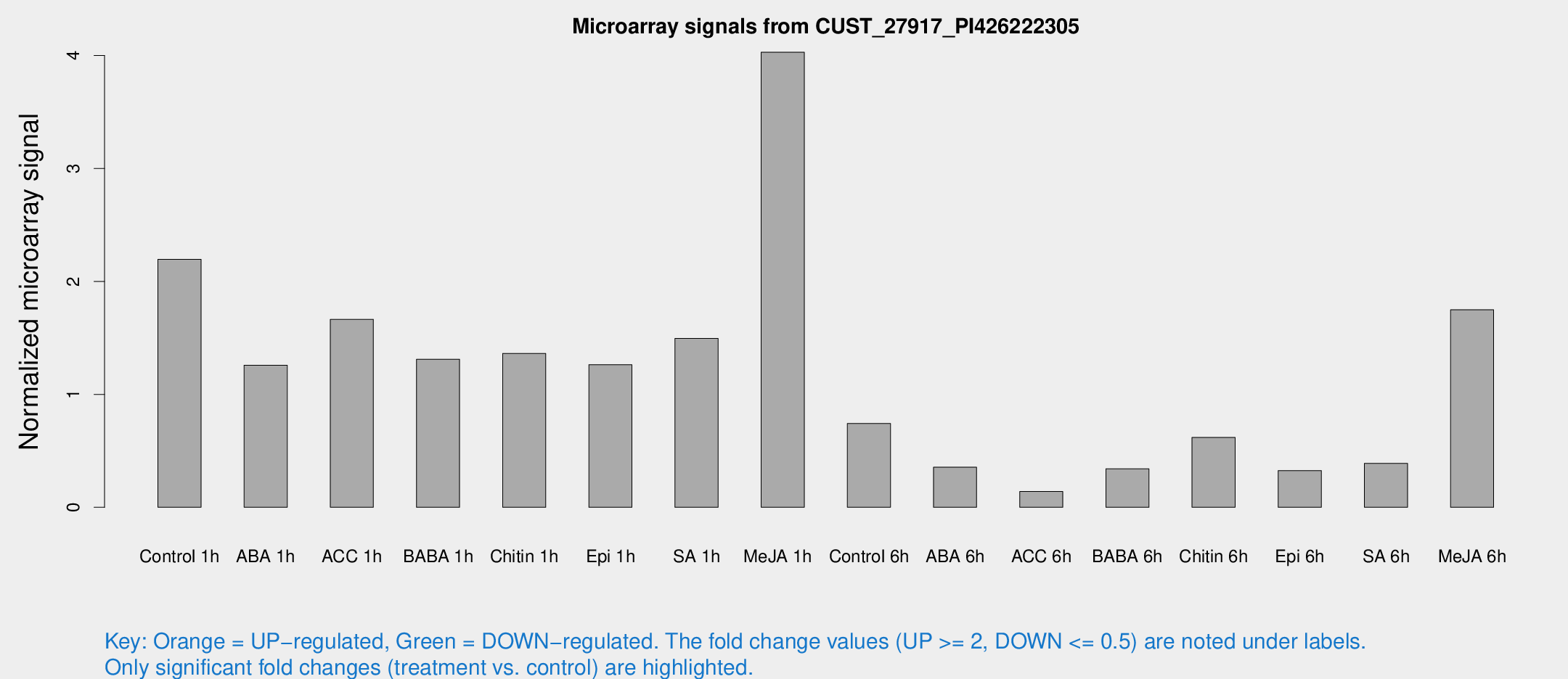
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1022.1 | 59.1295 | 2.195 | 0.171605 |
| ABA 1h | 543.293 | 107.632 | 1.25848 | 0.296389 |
| ACC 1h | 912.761 | 297.023 | 1.66365 | 0.581129 |
| BABA 1h | 589.728 | 34.3561 | 1.31087 | 0.144257 |
| Chitin 1h | 587.036 | 99.1607 | 1.36239 | 0.205969 |
| Epi 1h | 511.53 | 35.9863 | 1.26305 | 0.0862652 |
| SA 1h | 880.389 | 414.719 | 1.49523 | 0.802669 |
| Me-JA 1h | 1552.18 | 177.245 | 4.02924 | 0.232824 |
| Control 6h | 397.458 | 152.17 | 0.742036 | 0.23037 |
| ABA 6h | 183.024 | 38.8838 | 0.354846 | 0.0919061 |
| ACC 6h | 78.6393 | 16.6144 | 0.139835 | 0.0380386 |
| BABA 6h | 245.264 | 99.6642 | 0.340855 | 0.352856 |
| Chitin 6h | 309.568 | 31.0409 | 0.6181 | 0.0540677 |
| Epi 6h | 223.161 | 91.0824 | 0.3242 | 0.302972 |
| SA 6h | 185.803 | 32.644 | 0.389053 | 0.030432 |
| Me-JA 6h | 925.743 | 296.725 | 1.74883 | 0.75322 |
Source Transcript PGSC0003DMT400082027 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G52820.1 | +1 | 3e-26 | 106 | 48/117 (41%) | 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein | chr1:19669216-19670321 FORWARD LENGTH=317 |
