Probe CUST_28016_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28016_PI426222305 | JHI_St_60k_v1 | DMT400082026 | AACATCCCCTTGAGGTGCTGCCCTTTCTCTGTGAAAGAAATCACTAATGCTTTTTATGTG |
All Microarray Probes Designed to Gene DMG400032208
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_27917_PI426222305 | JHI_St_60k_v1 | DMT400082027 | TAACTTTAGGCATGGACAAATAATAGTTTGACATCTGCTGAACACAGAGTAGTACCAACA |
| CUST_28016_PI426222305 | JHI_St_60k_v1 | DMT400082026 | AACATCCCCTTGAGGTGCTGCCCTTTCTCTGTGAAAGAAATCACTAATGCTTTTTATGTG |
Microarray Signals from CUST_28016_PI426222305
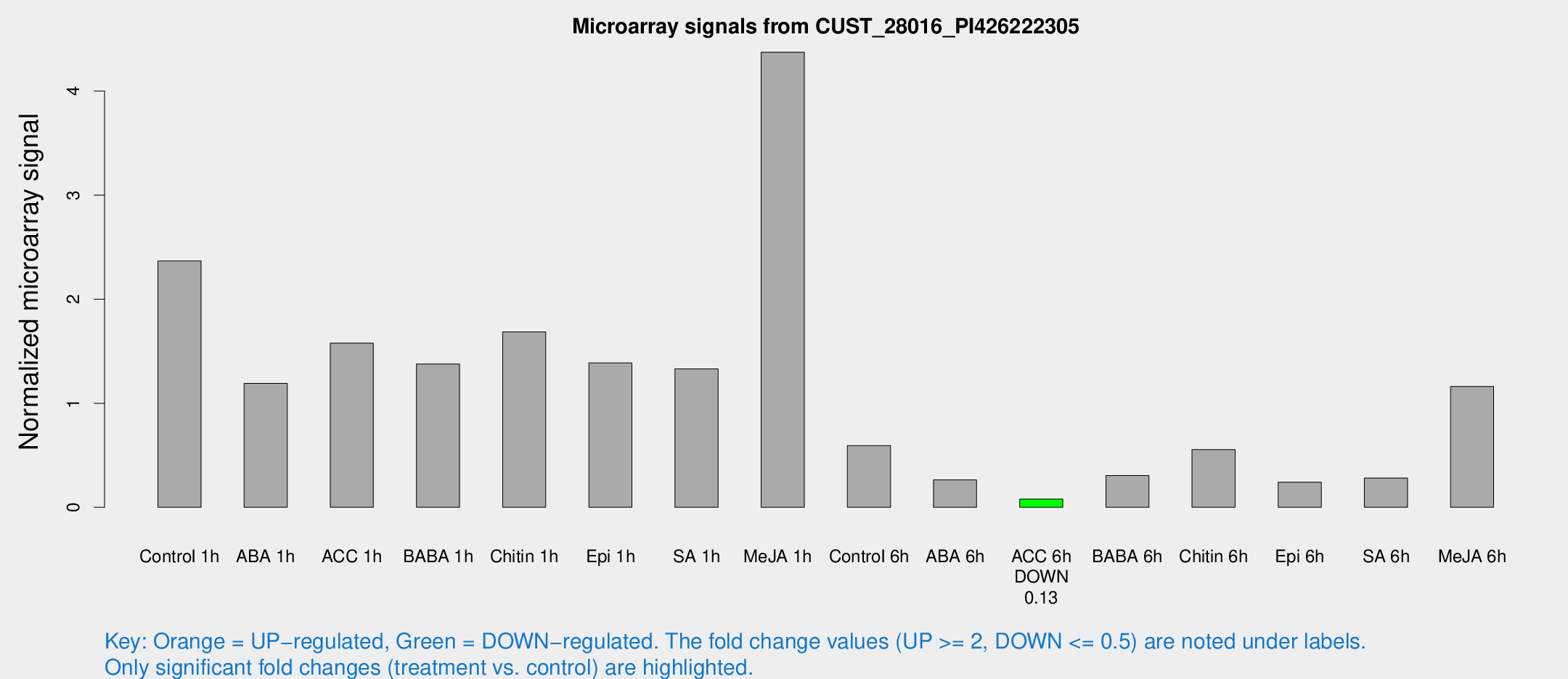
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1644.48 | 95.1424 | 2.3672 | 0.185988 |
| ABA 1h | 760.952 | 140.807 | 1.19108 | 0.254838 |
| ACC 1h | 1165.43 | 209.783 | 1.57729 | 0.187214 |
| BABA 1h | 928.109 | 79.0807 | 1.37739 | 0.079722 |
| Chitin 1h | 1092.72 | 222.974 | 1.68543 | 0.338794 |
| Epi 1h | 838.746 | 66.3424 | 1.38762 | 0.150612 |
| SA 1h | 1100.46 | 430.781 | 1.33009 | 0.612808 |
| Me-JA 1h | 2562.99 | 489.56 | 4.37358 | 0.446357 |
| Control 6h | 480.423 | 172.489 | 0.5924 | 0.203058 |
| ABA 6h | 195.536 | 12.7092 | 0.263944 | 0.0324688 |
| ACC 6h | 64.2451 | 9.404 | 0.0788393 | 0.0110367 |
| BABA 6h | 306.741 | 120.8 | 0.305003 | 0.245125 |
| Chitin 6h | 410.587 | 24.1354 | 0.554188 | 0.0494084 |
| Epi 6h | 255.388 | 106.69 | 0.241131 | 0.252274 |
| SA 6h | 202.869 | 47.4013 | 0.280835 | 0.0449153 |
| Me-JA 6h | 889.849 | 302.625 | 1.16065 | 0.301741 |
Source Transcript PGSC0003DMT400082026 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G52820.1 | +3 | 8e-10 | 45 | 22/44 (50%) | 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein | chr1:19669216-19670321 FORWARD LENGTH=317 |
