Probe CUST_26831_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_26831_PI426222305 | JHI_St_60k_v1 | DMT400034117 | GTTCATGGTTTGTCTGATTCGGAGTCTGAACTCCTTCGATATCGTGTTATCTCTCAGGTT |
All Microarray Probes Designed to Gene DMG400013108
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_26831_PI426222305 | JHI_St_60k_v1 | DMT400034117 | GTTCATGGTTTGTCTGATTCGGAGTCTGAACTCCTTCGATATCGTGTTATCTCTCAGGTT |
| CUST_26813_PI426222305 | JHI_St_60k_v1 | DMT400034118 | TTCTGCTTTCTAGTGTGCCTAGTATGTTTTGGACCATGTTGATTGAAAGCAACCTTGAAA |
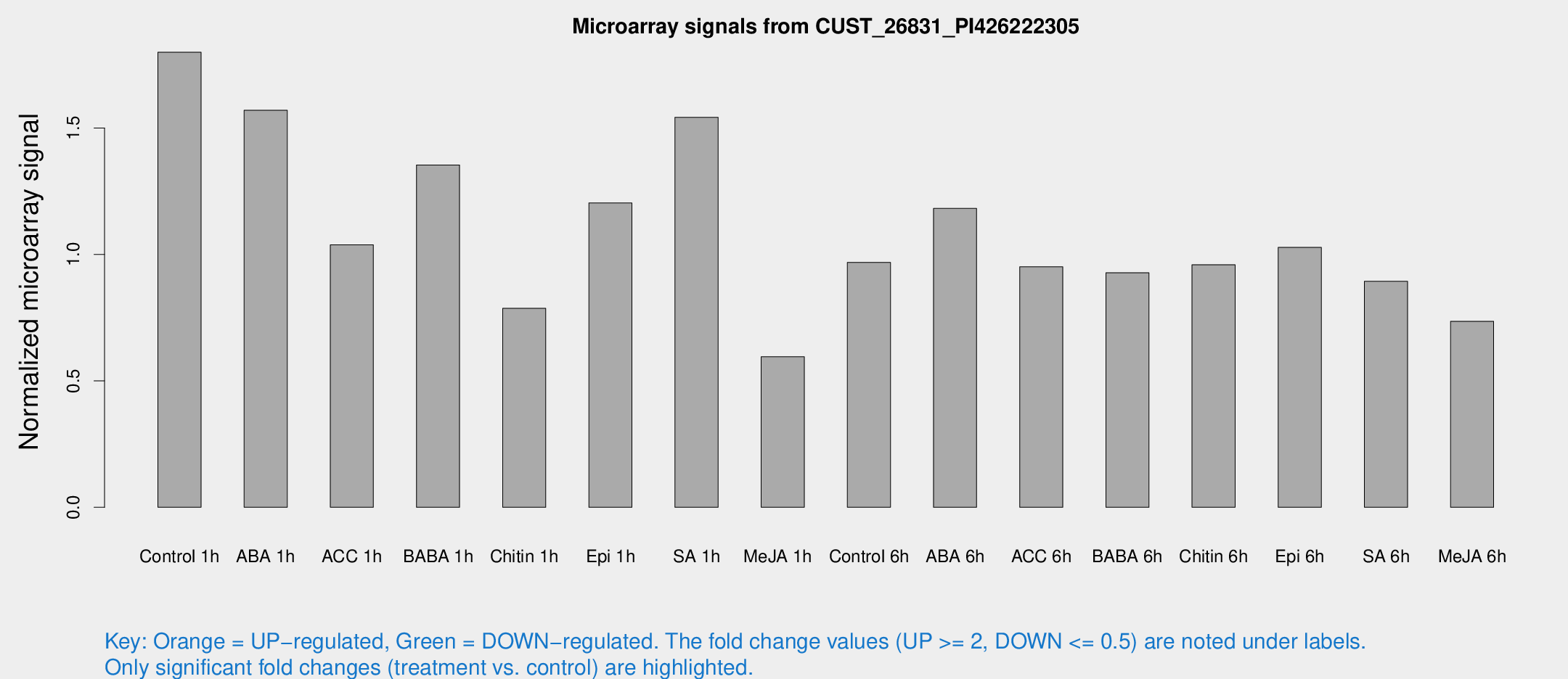
Microarray Signals from CUST_26831_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 6710.71 | 1155.85 | 1.79961 | 0.184256 |
| ABA 1h | 5519.98 | 1444.11 | 1.56982 | 0.440745 |
| ACC 1h | 5190.62 | 2043.66 | 1.03775 | 0.723088 |
| BABA 1h | 5226.81 | 1461.18 | 1.35378 | 0.324326 |
| Chitin 1h | 2703.29 | 646.121 | 0.78662 | 0.135079 |
| Epi 1h | 3823.31 | 400.576 | 1.20355 | 0.115395 |
| SA 1h | 6011.88 | 1301.87 | 1.54187 | 0.23577 |
| Me-JA 1h | 1851.97 | 428.656 | 0.595369 | 0.0968169 |
| Control 6h | 3690.44 | 759.445 | 0.968666 | 0.141959 |
| ABA 6h | 4677.59 | 709.253 | 1.18198 | 0.141248 |
| ACC 6h | 3992.56 | 369.626 | 0.95089 | 0.118419 |
| BABA 6h | 3799.89 | 351.481 | 0.927401 | 0.0559677 |
| Chitin 6h | 3729.37 | 320.042 | 0.95917 | 0.0602796 |
| Epi 6h | 4262.83 | 487.353 | 1.02771 | 0.0717545 |
| SA 6h | 3851.7 | 1389.49 | 0.893797 | 0.341536 |
| Me-JA 6h | 2817.9 | 640.685 | 0.735653 | 0.124044 |
Source Transcript PGSC0003DMT400034117 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G05300.1 | +1 | 3e-104 | 318 | 195/348 (56%) | zinc transporter 5 precursor | chr1:1545258-1547709 REVERSE LENGTH=360 |
