Probe CUST_26813_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_26813_PI426222305 | JHI_St_60k_v1 | DMT400034118 | TTCTGCTTTCTAGTGTGCCTAGTATGTTTTGGACCATGTTGATTGAAAGCAACCTTGAAA |
All Microarray Probes Designed to Gene DMG400013108
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_26831_PI426222305 | JHI_St_60k_v1 | DMT400034117 | GTTCATGGTTTGTCTGATTCGGAGTCTGAACTCCTTCGATATCGTGTTATCTCTCAGGTT |
| CUST_26813_PI426222305 | JHI_St_60k_v1 | DMT400034118 | TTCTGCTTTCTAGTGTGCCTAGTATGTTTTGGACCATGTTGATTGAAAGCAACCTTGAAA |
Microarray Signals from CUST_26813_PI426222305
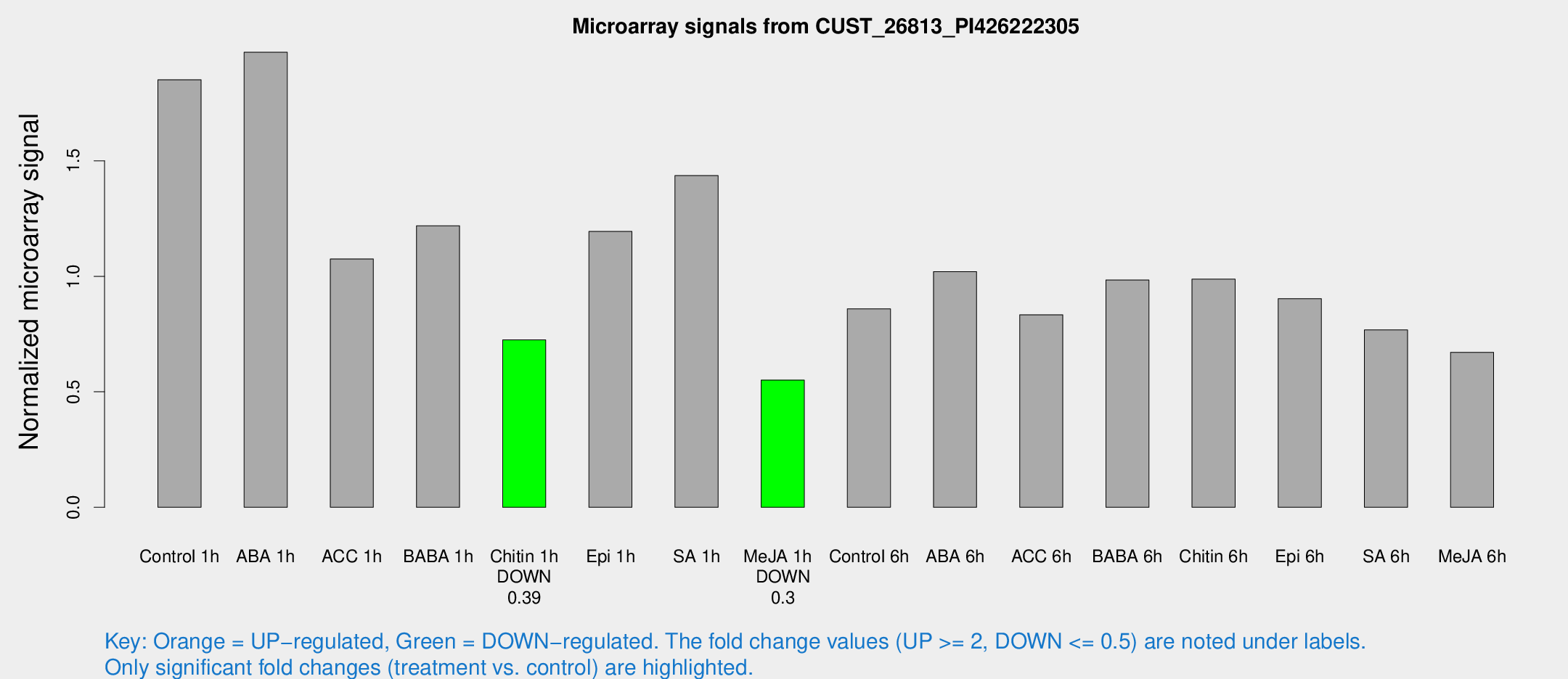
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 431.998 | 66.2441 | 1.85098 | 0.173089 |
| ABA 1h | 416.458 | 80.9731 | 1.96996 | 0.346108 |
| ACC 1h | 313.764 | 113.952 | 1.07515 | 0.560749 |
| BABA 1h | 276.192 | 45.1036 | 1.21882 | 0.0960174 |
| Chitin 1h | 150.262 | 15.0088 | 0.724653 | 0.0445209 |
| Epi 1h | 246.479 | 47.2476 | 1.19401 | 0.24414 |
| SA 1h | 342.851 | 46.5737 | 1.43604 | 0.14398 |
| Me-JA 1h | 109.733 | 29.7054 | 0.550819 | 0.10344 |
| Control 6h | 196.752 | 11.8083 | 0.859413 | 0.073863 |
| ABA 6h | 249.725 | 22.4379 | 1.02031 | 0.0605187 |
| ACC 6h | 220.483 | 21.0904 | 0.833493 | 0.108623 |
| BABA 6h | 253.182 | 20.2912 | 0.983942 | 0.0834496 |
| Chitin 6h | 241.026 | 14.9943 | 0.987834 | 0.0686902 |
| Epi 6h | 232.91 | 13.999 | 0.903167 | 0.0541963 |
| SA 6h | 194.655 | 56.1889 | 0.76818 | 0.197042 |
| Me-JA 6h | 161.646 | 37.5817 | 0.671086 | 0.108156 |
Source Transcript PGSC0003DMT400034118 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G05300.1 | +2 | 1e-26 | 105 | 68/102 (67%) | zinc transporter 5 precursor | chr1:1545258-1547709 REVERSE LENGTH=360 |
