Probe CUST_26281_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_26281_PI426222305 | JHI_St_60k_v1 | DMT400041601 | TTCGAGAGAAAGCAAAAGAAATGAGAGCCATATTTGGTAACAAAGAGCTTCATGATAGGT |
All Microarray Probes Designed to Gene DMG400016125
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_26281_PI426222305 | JHI_St_60k_v1 | DMT400041601 | TTCGAGAGAAAGCAAAAGAAATGAGAGCCATATTTGGTAACAAAGAGCTTCATGATAGGT |
| CUST_26259_PI426222305 | JHI_St_60k_v1 | DMT400041600 | TTCGAGAGAAAGCAAAAGAAATGAGAGCCATATTTGGTAACAAAGAGCTTCATGATAGGT |
Microarray Signals from CUST_26281_PI426222305
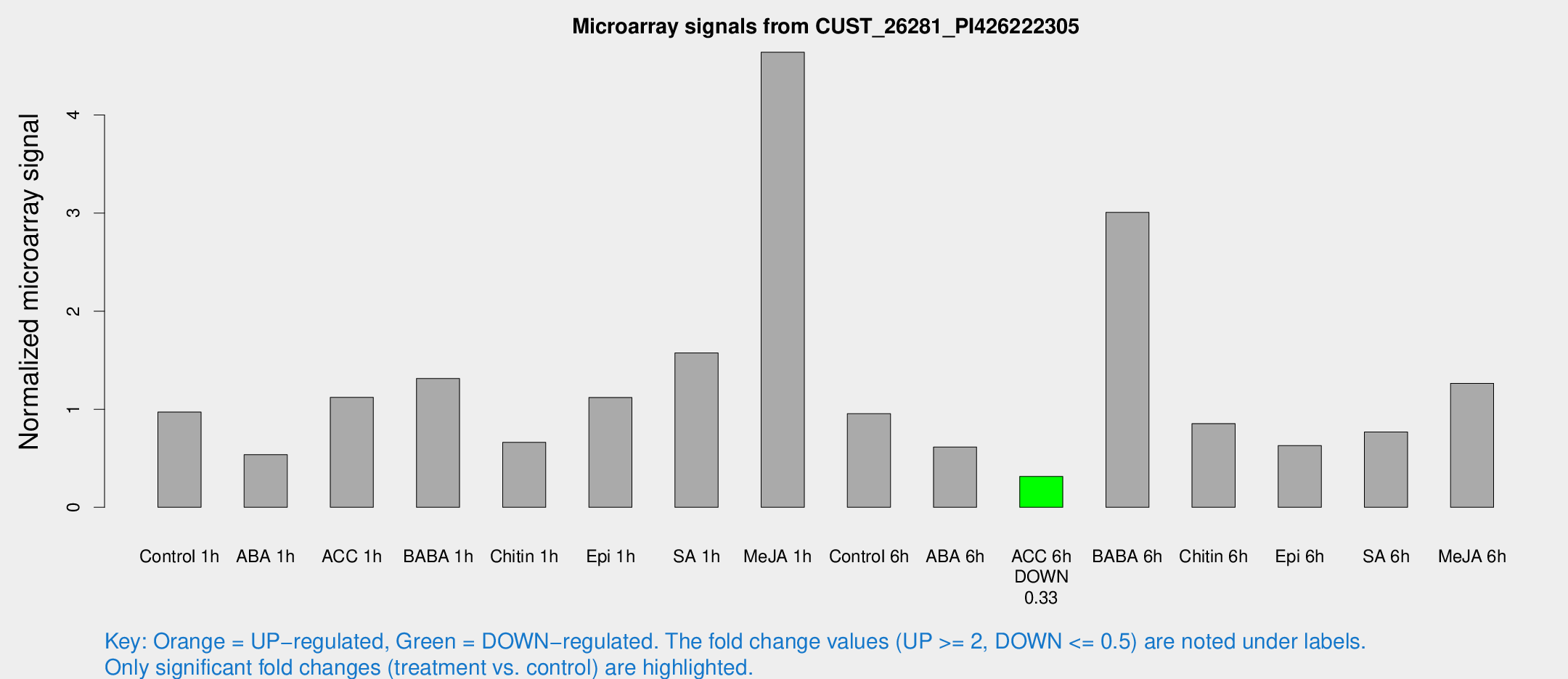
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 17.888 | 3.32478 | 0.971063 | 0.182155 |
| ABA 1h | 9.85628 | 4.00633 | 0.537312 | 0.267953 |
| ACC 1h | 21.6739 | 4.85223 | 1.12048 | 0.209429 |
| BABA 1h | 22.6662 | 3.41405 | 1.31284 | 0.213419 |
| Chitin 1h | 11.5682 | 3.5593 | 0.661956 | 0.211792 |
| Epi 1h | 17.4738 | 3.15771 | 1.11989 | 0.210609 |
| SA 1h | 32.2825 | 11.3429 | 1.57435 | 0.546401 |
| Me-JA 1h | 69.6668 | 12.1083 | 4.64053 | 0.676505 |
| Control 6h | 18.6171 | 4.97717 | 0.953972 | 0.218715 |
| ABA 6h | 12.3577 | 3.3399 | 0.614662 | 0.186865 |
| ACC 6h | 6.64093 | 3.81181 | 0.314427 | 0.177826 |
| BABA 6h | 435.362 | 421.394 | 3.00682 | 75.0347 |
| Chitin 6h | 16.9671 | 3.59703 | 0.853243 | 0.202741 |
| Epi 6h | 13.6181 | 3.68485 | 0.629518 | 0.235537 |
| SA 6h | 15.6485 | 4.89027 | 0.76826 | 0.25579 |
| Me-JA 6h | 23.1826 | 4.27963 | 1.26427 | 0.199332 |
Source Transcript PGSC0003DMT400041601 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G22590.1 | +1 | 3e-156 | 459 | 227/470 (48%) | UDP-Glycosyltransferase superfamily protein | chr2:9593012-9594424 FORWARD LENGTH=470 |
