Probe CUST_26259_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_26259_PI426222305 | JHI_St_60k_v1 | DMT400041600 | TTCGAGAGAAAGCAAAAGAAATGAGAGCCATATTTGGTAACAAAGAGCTTCATGATAGGT |
All Microarray Probes Designed to Gene DMG400016125
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_26281_PI426222305 | JHI_St_60k_v1 | DMT400041601 | TTCGAGAGAAAGCAAAAGAAATGAGAGCCATATTTGGTAACAAAGAGCTTCATGATAGGT |
| CUST_26259_PI426222305 | JHI_St_60k_v1 | DMT400041600 | TTCGAGAGAAAGCAAAAGAAATGAGAGCCATATTTGGTAACAAAGAGCTTCATGATAGGT |
Microarray Signals from CUST_26259_PI426222305
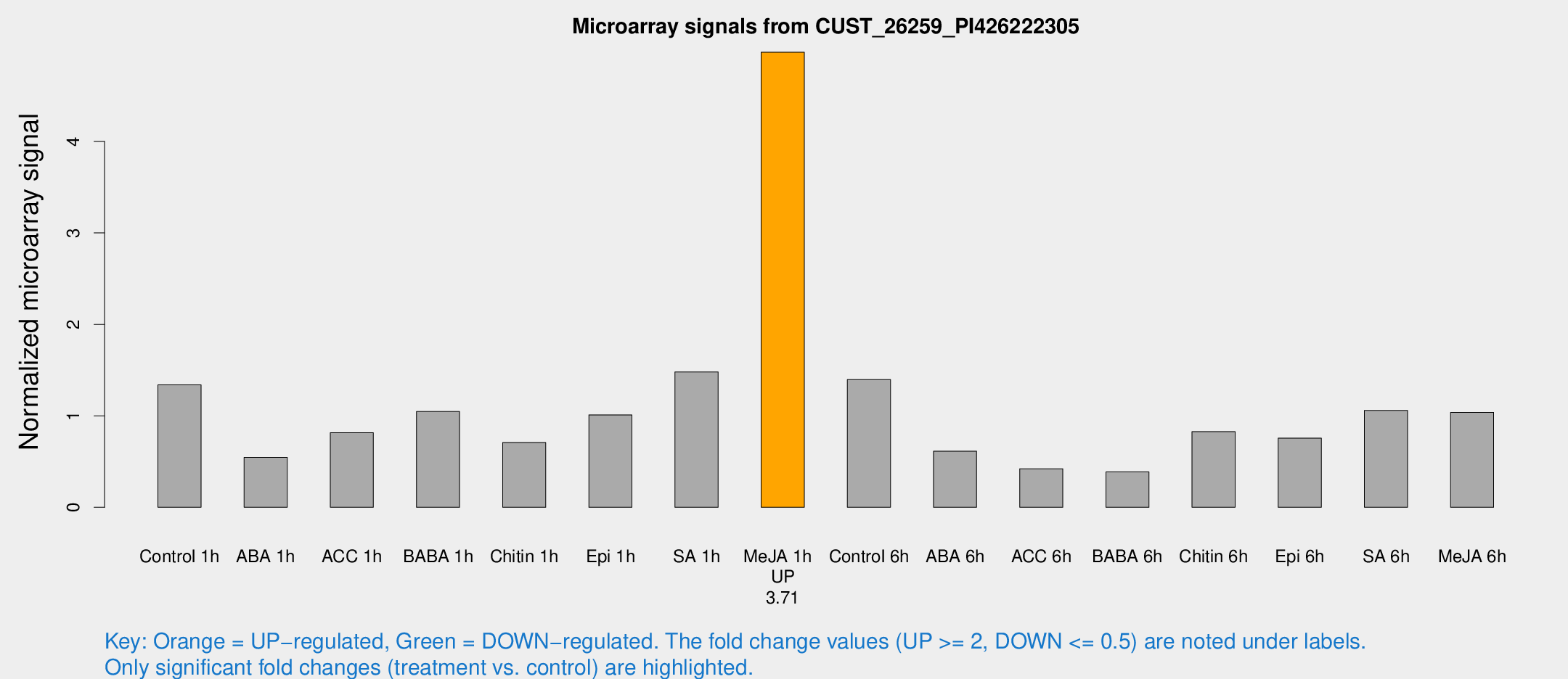
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 26.0359 | 3.6364 | 1.3394 | 0.187792 |
| ABA 1h | 11.4314 | 5.25485 | 0.54621 | 0.232134 |
| ACC 1h | 17.1986 | 3.80976 | 0.815494 | 0.261296 |
| BABA 1h | 20.2258 | 3.78645 | 1.04736 | 0.211247 |
| Chitin 1h | 14.5837 | 5.0639 | 0.707795 | 0.307599 |
| Epi 1h | 17.0483 | 3.50631 | 1.01008 | 0.211717 |
| SA 1h | 33.9041 | 13.1385 | 1.47942 | 0.575785 |
| Me-JA 1h | 80.2318 | 10.9282 | 4.97532 | 0.362903 |
| Control 6h | 27.2989 | 3.94953 | 1.39662 | 0.205514 |
| ABA 6h | 13.9297 | 3.81603 | 0.613802 | 0.225036 |
| ACC 6h | 9.5848 | 4.37608 | 0.421878 | 0.195521 |
| BABA 6h | 8.60881 | 4.06789 | 0.38631 | 0.193481 |
| Chitin 6h | 17.7726 | 4.07231 | 0.827844 | 0.209336 |
| Epi 6h | 17.0159 | 4.2423 | 0.756131 | 0.201893 |
| SA 6h | 20.7833 | 3.97293 | 1.05844 | 0.216917 |
| Me-JA 6h | 20.2279 | 3.81797 | 1.03817 | 0.199831 |
Source Transcript PGSC0003DMT400041600 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G22590.1 | +1 | 2e-157 | 459 | 227/470 (48%) | UDP-Glycosyltransferase superfamily protein | chr2:9593012-9594424 FORWARD LENGTH=470 |
