Probe CUST_2523_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_2523_PI426222305 | JHI_St_60k_v1 | DMT400028828 | TTTCTTTCTTCATTATCTTCCCGAAGCGTTAAGGGATGAGAACAAATGGCCCCGTCTTCC |
All Microarray Probes Designed to Gene DMG400011098
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_2355_PI426222305 | JHI_St_60k_v1 | DMT400028827 | CATCAGAACAAAAGGTCTTACTGGAAAATGTCAAATTGAATCACTTAAGTCCCCTAGATA |
| CUST_2523_PI426222305 | JHI_St_60k_v1 | DMT400028828 | TTTCTTTCTTCATTATCTTCCCGAAGCGTTAAGGGATGAGAACAAATGGCCCCGTCTTCC |
Microarray Signals from CUST_2523_PI426222305
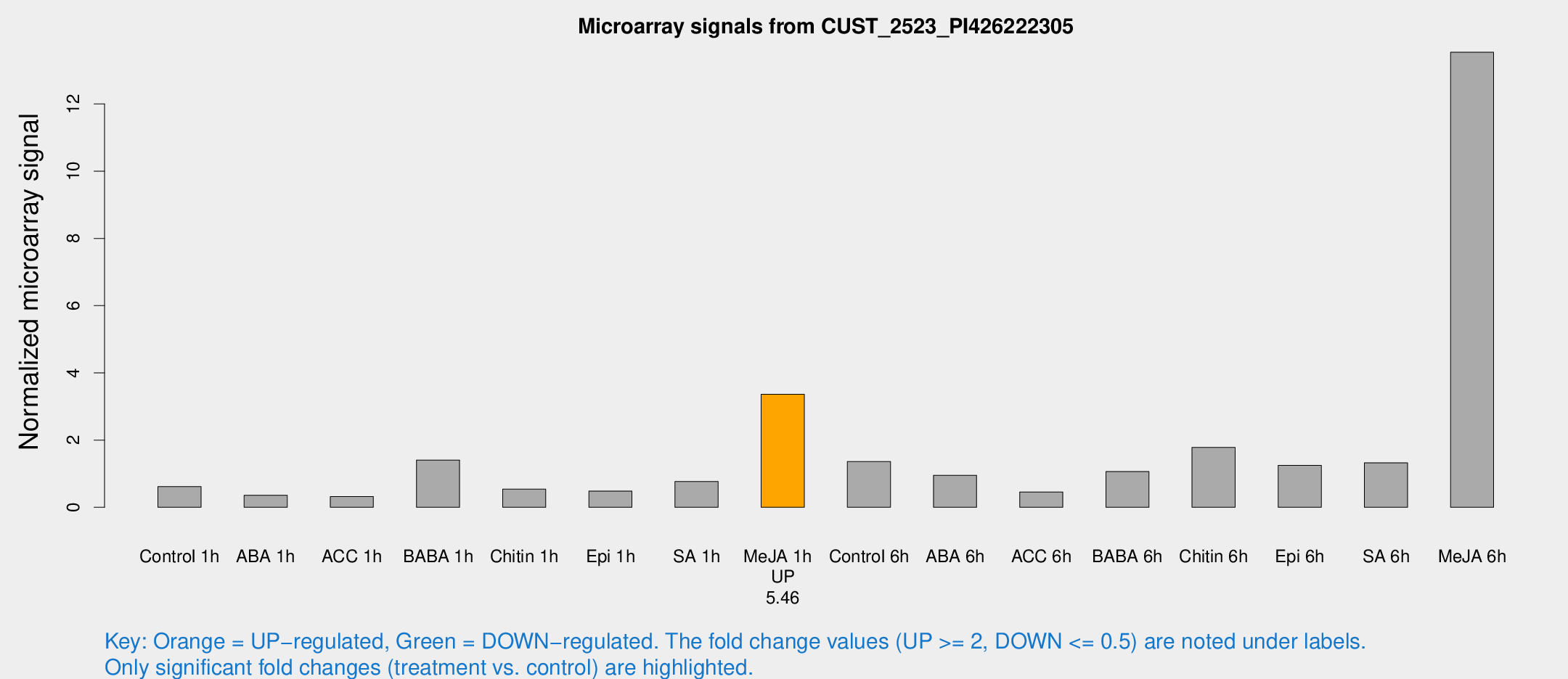
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 20.7447 | 7.21945 | 0.615876 | 0.258825 |
| ABA 1h | 11.7823 | 5.59237 | 0.358475 | 0.160202 |
| ACC 1h | 10.2781 | 4.00201 | 0.317954 | 0.136859 |
| BABA 1h | 43.3283 | 10.5148 | 1.407 | 0.516378 |
| Chitin 1h | 15.639 | 3.9285 | 0.541252 | 0.19207 |
| Epi 1h | 14.4419 | 5.09903 | 0.482681 | 0.215836 |
| SA 1h | 32.9978 | 15.6298 | 0.766388 | 0.601505 |
| Me-JA 1h | 86.1655 | 15.7516 | 3.36143 | 0.432075 |
| Control 6h | 42.7016 | 7.53068 | 1.36115 | 0.152205 |
| ABA 6h | 30.7947 | 4.24991 | 0.953222 | 0.133729 |
| ACC 6h | 19.5486 | 8.60373 | 0.456195 | 0.189384 |
| BABA 6h | 54.2372 | 27.1264 | 1.06346 | 1.29612 |
| Chitin 6h | 57.3582 | 5.5847 | 1.78088 | 0.172959 |
| Epi 6h | 52.9301 | 19.3221 | 1.24725 | 0.964674 |
| SA 6h | 42.2141 | 10.2854 | 1.32178 | 0.211818 |
| Me-JA 6h | 457.185 | 132.309 | 13.5387 | 6.98642 |
Source Transcript PGSC0003DMT400028828 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G55970.1 | +3 | 3e-147 | 426 | 209/366 (57%) | jasmonate-regulated gene 21 | chr3:20766970-20769264 REVERSE LENGTH=363 |
