Probe CUST_25234_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_25234_PI426222305 | JHI_St_60k_v1 | DMT400014953 | GGTAGAGACCATTTTTATATTGTGGGCAGTGTAAATGGTTAATTTTGAAGATGTGTAGCT |
All Microarray Probes Designed to Gene DMG400005837
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_25096_PI426222305 | JHI_St_60k_v1 | DMT400014952 | TTATTCCTTAGCAGCAATGCCTTCTTTTGATCCTGAGTTGATATGGGCAGTTCTGGCTAG |
| CUST_25234_PI426222305 | JHI_St_60k_v1 | DMT400014953 | GGTAGAGACCATTTTTATATTGTGGGCAGTGTAAATGGTTAATTTTGAAGATGTGTAGCT |
Microarray Signals from CUST_25234_PI426222305
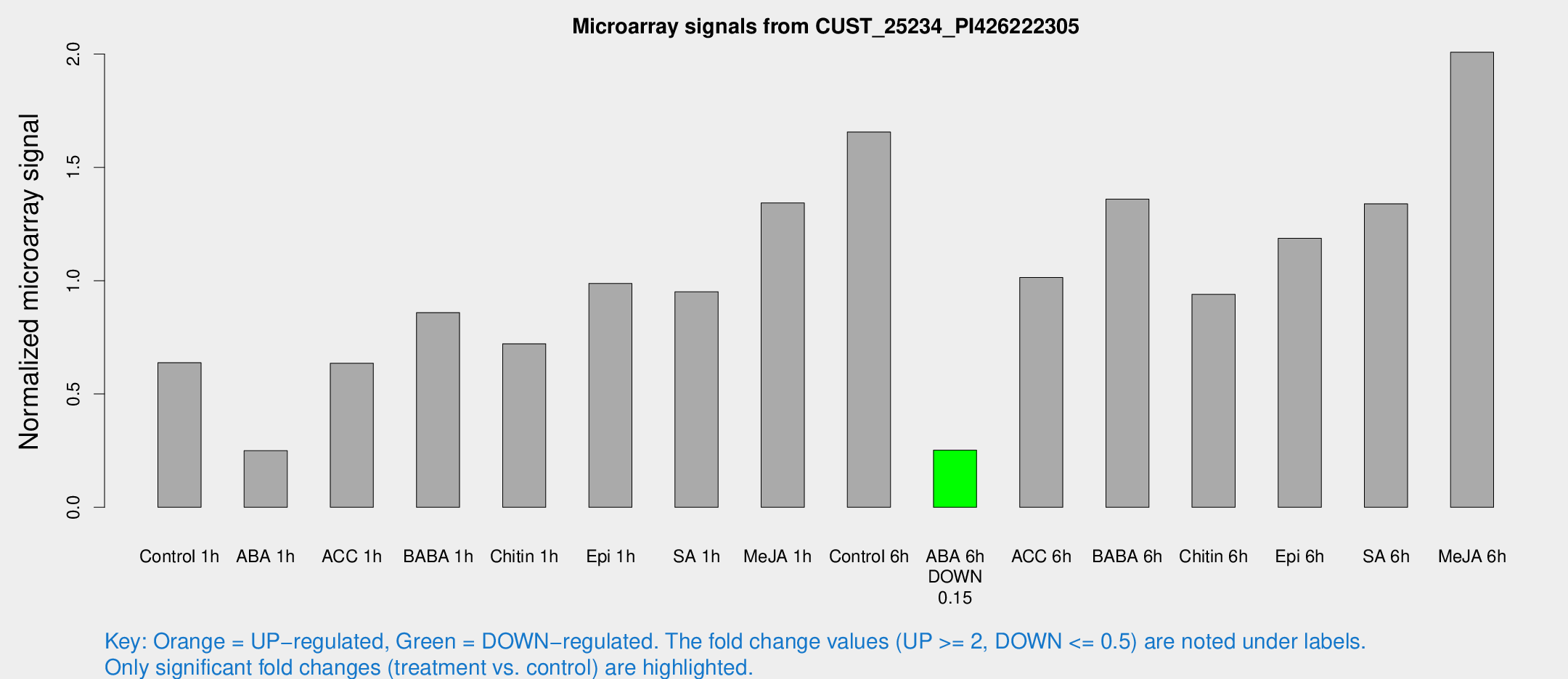
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 448.374 | 78.5461 | 0.638134 | 0.133381 |
| ABA 1h | 158.343 | 37.4016 | 0.249931 | 0.0510713 |
| ACC 1h | 446.834 | 44.3278 | 0.635654 | 0.0384374 |
| BABA 1h | 587.366 | 116.621 | 0.858968 | 0.103185 |
| Chitin 1h | 446.598 | 53.5126 | 0.721526 | 0.14287 |
| Epi 1h | 592.845 | 84.7618 | 0.987615 | 0.138692 |
| SA 1h | 663.319 | 38.4923 | 0.951203 | 0.0836007 |
| Me-JA 1h | 749.522 | 69.2216 | 1.34323 | 0.108128 |
| Control 6h | 1184.4 | 242.699 | 1.65617 | 0.238458 |
| ABA 6h | 184.766 | 25.0946 | 0.252019 | 0.0223265 |
| ACC 6h | 801.166 | 104.227 | 1.01415 | 0.0587473 |
| BABA 6h | 1070.6 | 196.612 | 1.35973 | 0.216673 |
| Chitin 6h | 681.465 | 50.0408 | 0.939628 | 0.0544864 |
| Epi 6h | 936.076 | 170.902 | 1.18689 | 0.246642 |
| SA 6h | 946.352 | 197.845 | 1.33934 | 0.163452 |
| Me-JA 6h | 1454.72 | 362.674 | 2.00792 | 0.458767 |
Source Transcript PGSC0003DMT400014953 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G64380.1 | +2 | 1e-50 | 178 | 149/314 (47%) | Integrase-type DNA-binding superfamily protein | chr1:23890981-23891988 REVERSE LENGTH=335 |
