Probe CUST_25096_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_25096_PI426222305 | JHI_St_60k_v1 | DMT400014952 | TTATTCCTTAGCAGCAATGCCTTCTTTTGATCCTGAGTTGATATGGGCAGTTCTGGCTAG |
All Microarray Probes Designed to Gene DMG400005837
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_25096_PI426222305 | JHI_St_60k_v1 | DMT400014952 | TTATTCCTTAGCAGCAATGCCTTCTTTTGATCCTGAGTTGATATGGGCAGTTCTGGCTAG |
| CUST_25234_PI426222305 | JHI_St_60k_v1 | DMT400014953 | GGTAGAGACCATTTTTATATTGTGGGCAGTGTAAATGGTTAATTTTGAAGATGTGTAGCT |
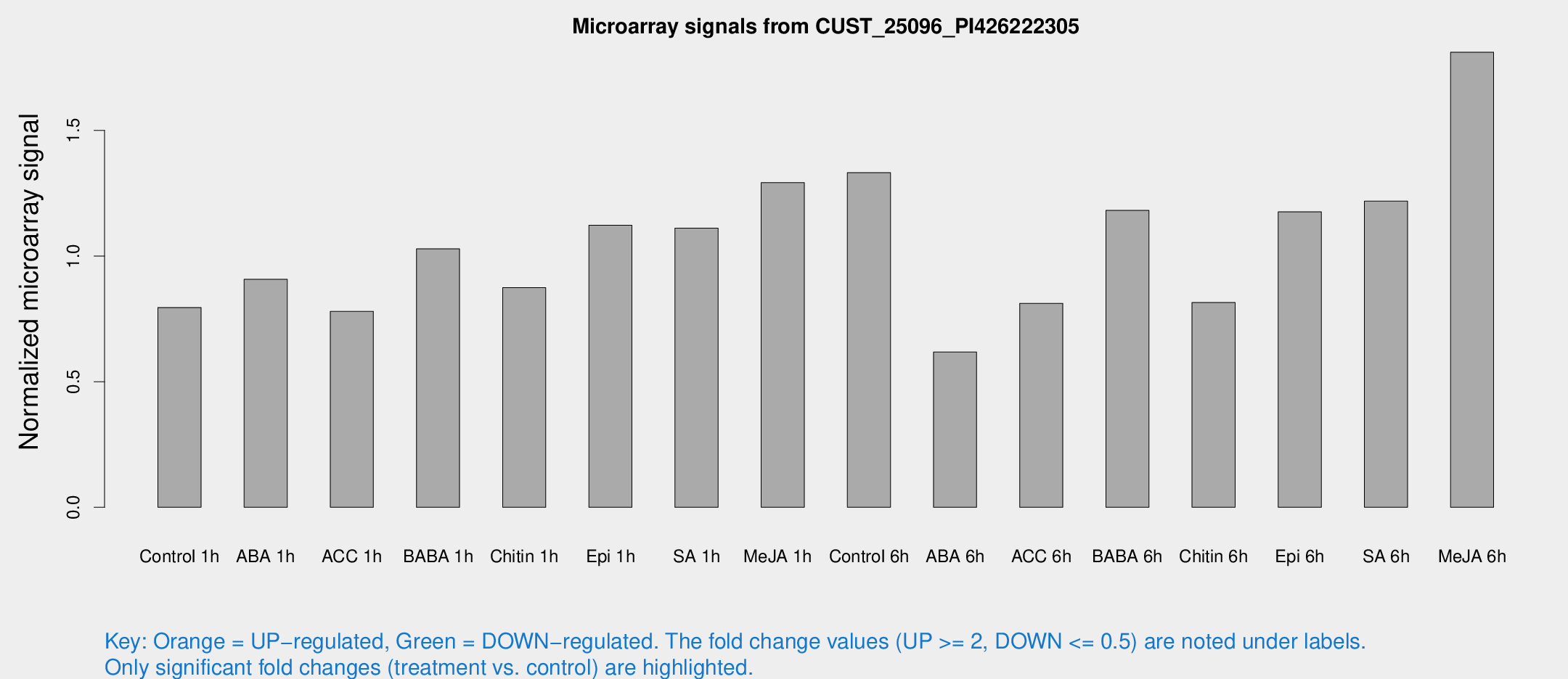
Microarray Signals from CUST_25096_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 780.096 | 94.7282 | 0.795347 | 0.083839 |
| ABA 1h | 795.599 | 118.256 | 0.90757 | 0.0995345 |
| ACC 1h | 779.87 | 67.7617 | 0.779835 | 0.0451946 |
| BABA 1h | 1019.48 | 232.79 | 1.02868 | 0.16446 |
| Chitin 1h | 765.141 | 63.1848 | 0.874524 | 0.0720546 |
| Epi 1h | 953.145 | 113.275 | 1.12264 | 0.116517 |
| SA 1h | 1117.38 | 131.741 | 1.11086 | 0.0671639 |
| Me-JA 1h | 1034.34 | 128.625 | 1.29211 | 0.0920325 |
| Control 6h | 1395.9 | 342.913 | 1.33193 | 0.279172 |
| ABA 6h | 644.894 | 80.5963 | 0.618076 | 0.0422856 |
| ACC 6h | 919.854 | 141.243 | 0.811774 | 0.0470181 |
| BABA 6h | 1297.12 | 152.363 | 1.18183 | 0.0964856 |
| Chitin 6h | 854.837 | 119.894 | 0.814951 | 0.0882865 |
| Epi 6h | 1311.59 | 207.637 | 1.176 | 0.200968 |
| SA 6h | 1259.04 | 326.212 | 1.21866 | 0.221879 |
| Me-JA 6h | 1844.19 | 430.546 | 1.81136 | 0.372106 |
Source Transcript PGSC0003DMT400014952 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G64380.1 | +1 | 4e-30 | 114 | 99/192 (52%) | Integrase-type DNA-binding superfamily protein | chr1:23890981-23891988 REVERSE LENGTH=335 |
