Probe CUST_25021_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_25021_PI426222305 | JHI_St_60k_v1 | DMT400074840 | TTTTTATTGTTGGACACAGATTCGATTCTACTGTTTGCAAAATCGAGGAAGCGGCGCGTC |
All Microarray Probes Designed to Gene DMG400029105
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_25029_PI426222305 | JHI_St_60k_v1 | DMT400074839 | TGAAGCTTGTTACACCCTTGGCATGATTCGATTCTACTGTTTGCAAAATCGAGGAAGCGG |
| CUST_25021_PI426222305 | JHI_St_60k_v1 | DMT400074840 | TTTTTATTGTTGGACACAGATTCGATTCTACTGTTTGCAAAATCGAGGAAGCGGCGCGTC |
Microarray Signals from CUST_25021_PI426222305
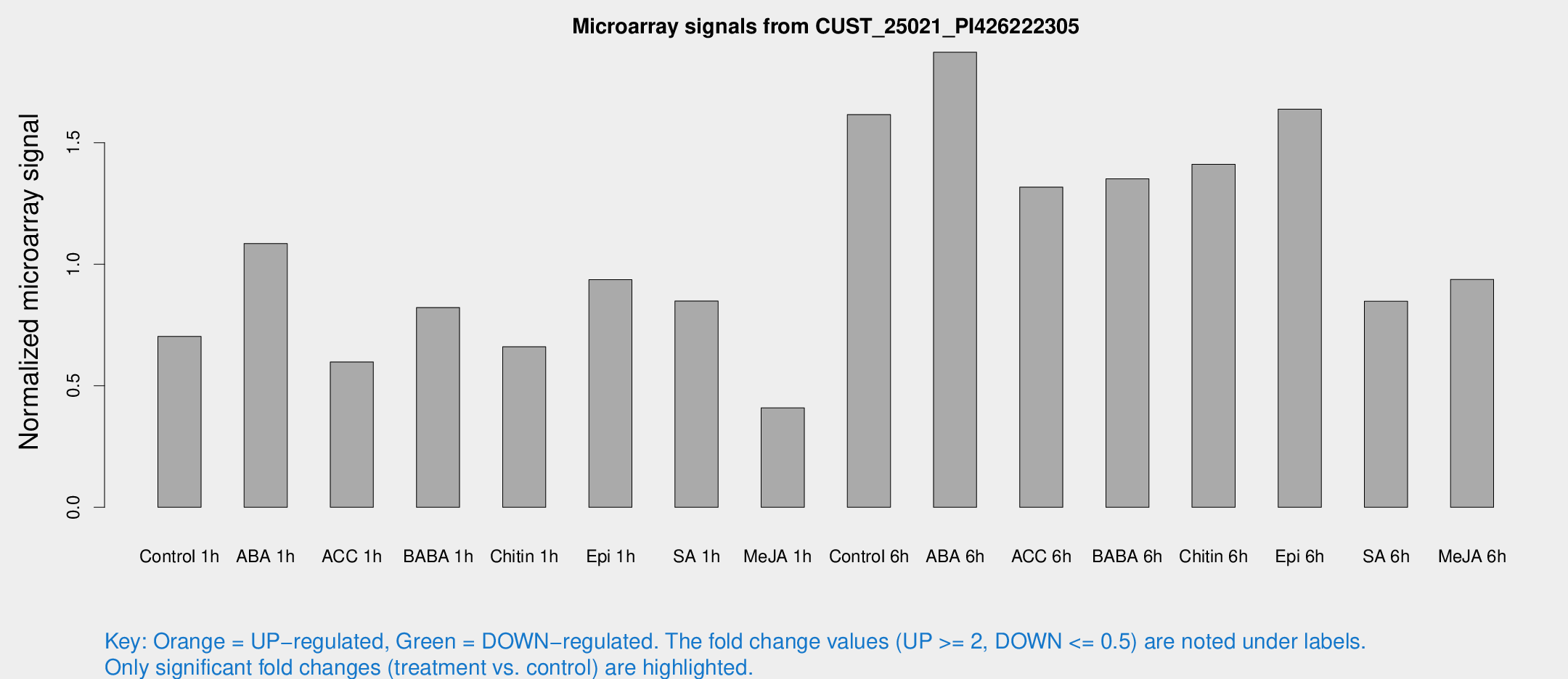
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 138.104 | 13.431 | 0.703115 | 0.0436722 |
| ABA 1h | 193.42 | 37.7525 | 1.08541 | 0.130278 |
| ACC 1h | 133.651 | 38.1926 | 0.597907 | 0.178786 |
| BABA 1h | 163.943 | 37.1332 | 0.821742 | 0.131651 |
| Chitin 1h | 115.814 | 7.43034 | 0.660389 | 0.0422232 |
| Epi 1h | 161.515 | 24.8259 | 0.936374 | 0.128508 |
| SA 1h | 176.411 | 36.7305 | 0.848651 | 0.130796 |
| Me-JA 1h | 65.6447 | 6.70845 | 0.409288 | 0.0352715 |
| Control 6h | 335.183 | 83.8224 | 1.61564 | 0.315894 |
| ABA 6h | 406.969 | 84.4772 | 1.87222 | 0.382181 |
| ACC 6h | 308.676 | 71.3351 | 1.31725 | 0.105444 |
| BABA 6h | 305.374 | 57.5492 | 1.35113 | 0.23339 |
| Chitin 6h | 305.728 | 61.4277 | 1.41115 | 0.309238 |
| Epi 6h | 376.285 | 84.3182 | 1.6382 | 0.393923 |
| SA 6h | 171.4 | 36.9474 | 0.847508 | 0.111083 |
| Me-JA 6h | 210.368 | 67.0062 | 0.937667 | 0.34444 |
Source Transcript PGSC0003DMT400074840 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G67340.1 | +2 | 5e-127 | 360 | 176/231 (76%) | HCP-like superfamily protein with MYND-type zinc finger | chr1:25230323-25231622 FORWARD LENGTH=379 |
