Probe CUST_25029_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_25029_PI426222305 | JHI_St_60k_v1 | DMT400074839 | TGAAGCTTGTTACACCCTTGGCATGATTCGATTCTACTGTTTGCAAAATCGAGGAAGCGG |
All Microarray Probes Designed to Gene DMG400029105
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_25029_PI426222305 | JHI_St_60k_v1 | DMT400074839 | TGAAGCTTGTTACACCCTTGGCATGATTCGATTCTACTGTTTGCAAAATCGAGGAAGCGG |
| CUST_25021_PI426222305 | JHI_St_60k_v1 | DMT400074840 | TTTTTATTGTTGGACACAGATTCGATTCTACTGTTTGCAAAATCGAGGAAGCGGCGCGTC |
Microarray Signals from CUST_25029_PI426222305
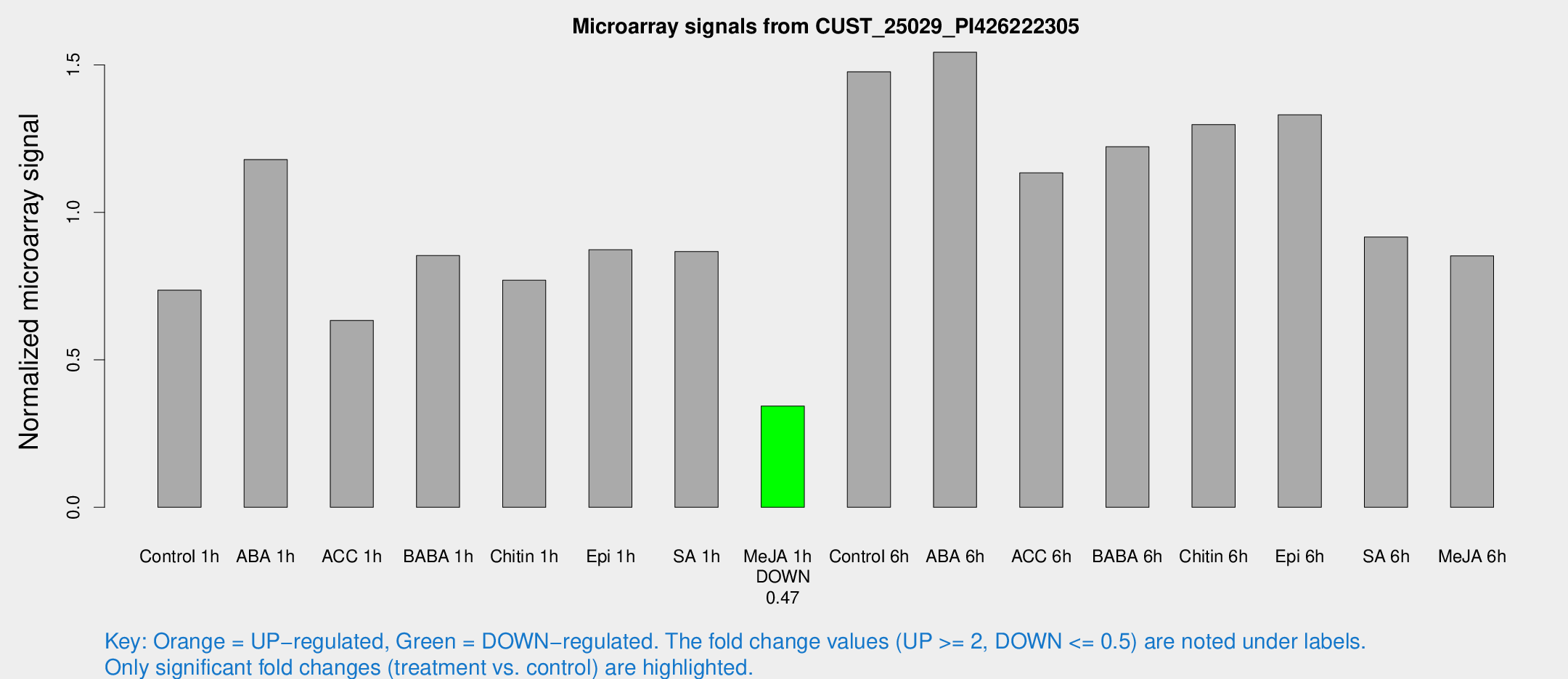
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 643.241 | 61.3539 | 0.736158 | 0.0426656 |
| ABA 1h | 928.617 | 154.498 | 1.17902 | 0.113511 |
| ACC 1h | 611.5 | 152.616 | 0.633674 | 0.147011 |
| BABA 1h | 779.792 | 207.609 | 0.853432 | 0.188535 |
| Chitin 1h | 603.018 | 45.6048 | 0.769954 | 0.044641 |
| Epi 1h | 671.095 | 105.214 | 0.873209 | 0.118143 |
| SA 1h | 807.674 | 181.344 | 0.866937 | 0.141967 |
| Me-JA 1h | 249.767 | 43.0726 | 0.343285 | 0.0286313 |
| Control 6h | 1375.72 | 353.277 | 1.47664 | 0.302546 |
| ABA 6h | 1462.06 | 230.951 | 1.54285 | 0.199744 |
| ACC 6h | 1195.44 | 274.331 | 1.13398 | 0.174122 |
| BABA 6h | 1231.67 | 223.591 | 1.22257 | 0.209292 |
| Chitin 6h | 1215.91 | 150.636 | 1.29776 | 0.154419 |
| Epi 6h | 1342.96 | 253.271 | 1.33085 | 0.252682 |
| SA 6h | 843.989 | 202.786 | 0.916243 | 0.155601 |
| Me-JA 6h | 820.392 | 233.336 | 0.852611 | 0.2462 |
Source Transcript PGSC0003DMT400074839 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G67340.1 | +1 | 5e-167 | 483 | 237/337 (70%) | HCP-like superfamily protein with MYND-type zinc finger | chr1:25230323-25231622 FORWARD LENGTH=379 |
