Probe CUST_2500_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_2500_PI426222305 | JHI_St_60k_v1 | DMT400027149 | CAGTAGCAACATTCACACAATAGGACTGAAATACTTAGATGTTATGTGGAAGCTAACATT |
All Microarray Probes Designed to Gene DMG400010470
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_2500_PI426222305 | JHI_St_60k_v1 | DMT400027149 | CAGTAGCAACATTCACACAATAGGACTGAAATACTTAGATGTTATGTGGAAGCTAACATT |
| CUST_2086_PI426222305 | JHI_St_60k_v1 | DMT400027148 | ACCGTAAGAAGTCCTATAGTAGTTGCTCCAGCGCTTGATGCCCCAAATGAAAATGATTGA |
Microarray Signals from CUST_2500_PI426222305
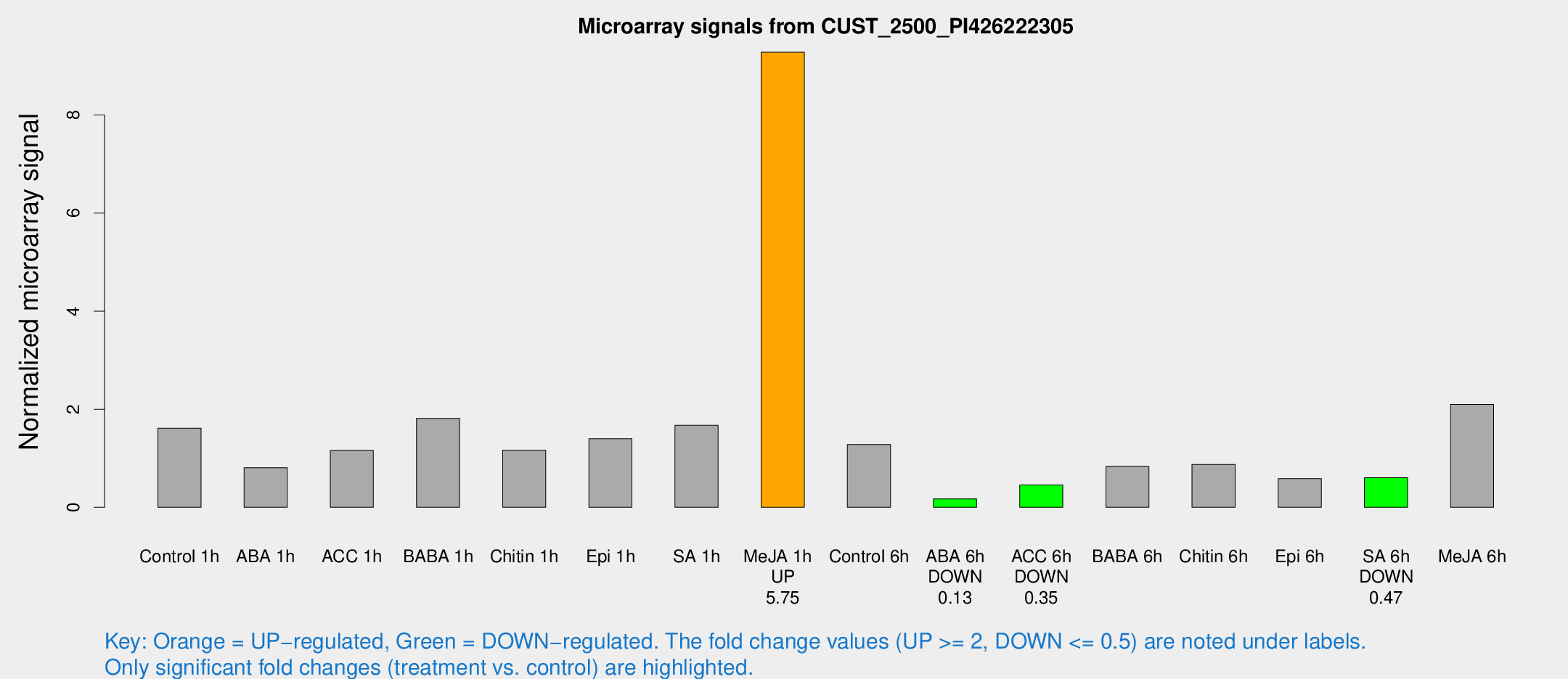
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 675.328 | 201.913 | 1.61442 | 0.496963 |
| ABA 1h | 313.368 | 122.128 | 0.808432 | 0.260179 |
| ACC 1h | 507.725 | 178.608 | 1.16223 | 0.362169 |
| BABA 1h | 691.929 | 128.639 | 1.81379 | 0.19 |
| Chitin 1h | 402.295 | 39.4031 | 1.16628 | 0.207275 |
| Epi 1h | 479.517 | 90.9438 | 1.39855 | 0.273966 |
| SA 1h | 682.163 | 129.7 | 1.67298 | 0.483847 |
| Me-JA 1h | 2916.32 | 323.801 | 9.28164 | 0.536013 |
| Control 6h | 521.099 | 133.071 | 1.28233 | 0.226245 |
| ABA 6h | 77.3244 | 22.2287 | 0.172327 | 0.0679118 |
| ACC 6h | 200.517 | 22.1899 | 0.454465 | 0.0281196 |
| BABA 6h | 365.139 | 63.0765 | 0.834126 | 0.127629 |
| Chitin 6h | 356.363 | 29.2154 | 0.875153 | 0.0809181 |
| Epi 6h | 264.857 | 55.8383 | 0.586205 | 0.186379 |
| SA 6h | 233.904 | 41.3422 | 0.604181 | 0.0769837 |
| Me-JA 6h | 809.228 | 112.167 | 2.09979 | 0.346444 |
Source Transcript PGSC0003DMT400027149 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G67090.1 | +1 | 2e-175 | 531 | 311/728 (43%) | Subtilisin-like serine endopeptidase family protein | chr5:26774111-26776321 REVERSE LENGTH=736 |
