Probe CUST_24974_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24974_PI426222305 | JHI_St_60k_v1 | DMT400024087 | CACTTGGAATATCTGCAACTAGAAAGAATGATCTTCACTTGATTGCCATTTCACATGATT |
All Microarray Probes Designed to Gene DMG400009315
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24842_PI426222305 | JHI_St_60k_v1 | DMT400024086 | CACTTGGAATATCTGCAACTAGAAAGAATGATCTTCACTTGATTGCCATTTCACATGATT |
| CUST_24974_PI426222305 | JHI_St_60k_v1 | DMT400024087 | CACTTGGAATATCTGCAACTAGAAAGAATGATCTTCACTTGATTGCCATTTCACATGATT |
Microarray Signals from CUST_24974_PI426222305
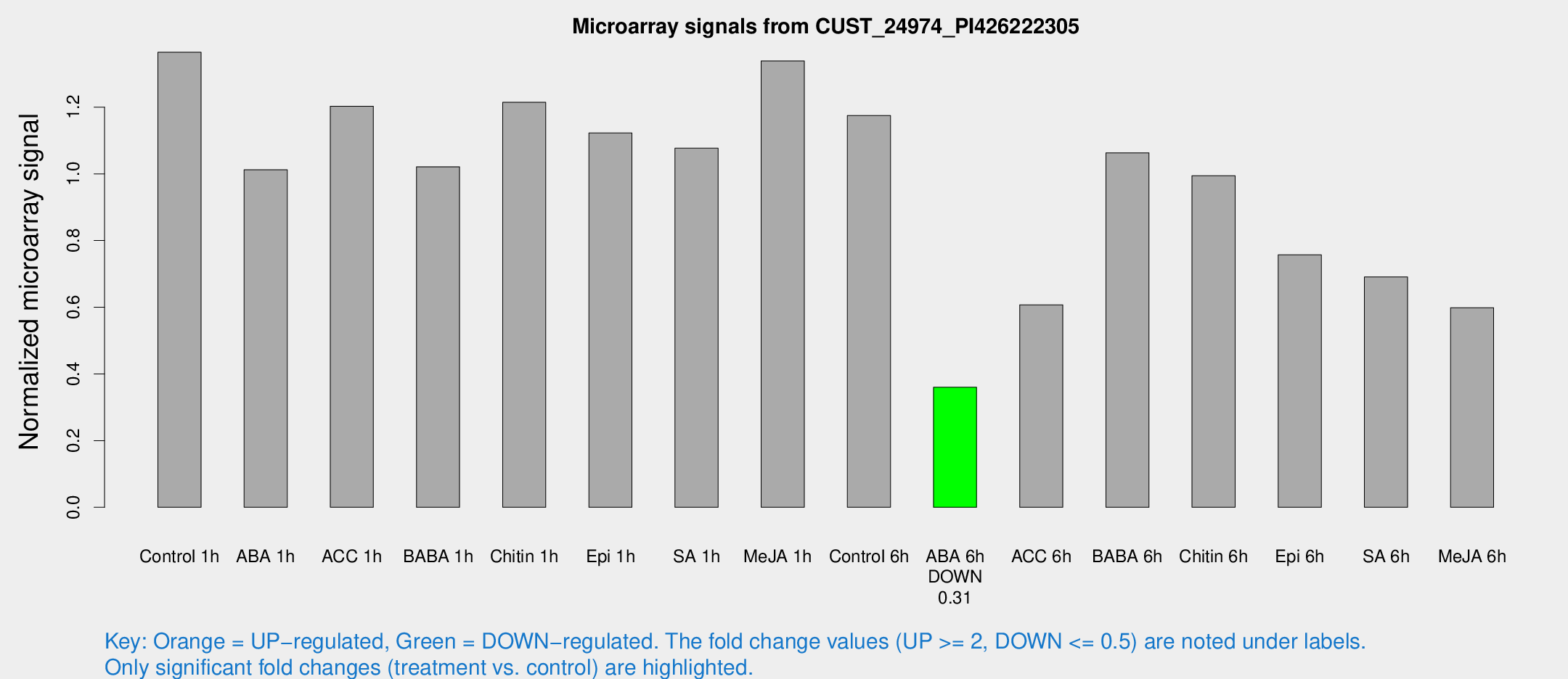
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1072.7 | 158.833 | 1.36465 | 0.109852 |
| ABA 1h | 711.101 | 132.529 | 1.01207 | 0.114839 |
| ACC 1h | 962.925 | 113.978 | 1.20281 | 0.115325 |
| BABA 1h | 793.164 | 155.646 | 1.0212 | 0.122288 |
| Chitin 1h | 858.084 | 125.041 | 1.21444 | 0.159316 |
| Epi 1h | 758.126 | 87.7563 | 1.12248 | 0.11472 |
| SA 1h | 861.546 | 92.5833 | 1.0768 | 0.0622929 |
| Me-JA 1h | 844.603 | 59.1987 | 1.33853 | 0.0774404 |
| Control 6h | 1033.76 | 319.895 | 1.17493 | 0.379535 |
| ABA 6h | 298.26 | 33.635 | 0.360086 | 0.0337544 |
| ACC 6h | 560.115 | 121.176 | 0.607291 | 0.0444303 |
| BABA 6h | 918.858 | 57.6156 | 1.06302 | 0.0767731 |
| Chitin 6h | 817.068 | 48.9003 | 0.994366 | 0.0575731 |
| Epi 6h | 666.805 | 83.6285 | 0.757397 | 0.0924095 |
| SA 6h | 534.819 | 67.547 | 0.691038 | 0.0401652 |
| Me-JA 6h | 473.624 | 79.1321 | 0.598655 | 0.0645285 |
Source Transcript PGSC0003DMT400024087 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G26320.1 | +3 | 6e-104 | 256 | 135/336 (40%) | cytochrome P450, family 71, subfamily B, polypeptide 36 | chr3:9644383-9646064 REVERSE LENGTH=500 |
