Probe CUST_24842_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24842_PI426222305 | JHI_St_60k_v1 | DMT400024086 | CACTTGGAATATCTGCAACTAGAAAGAATGATCTTCACTTGATTGCCATTTCACATGATT |
All Microarray Probes Designed to Gene DMG400009315
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24842_PI426222305 | JHI_St_60k_v1 | DMT400024086 | CACTTGGAATATCTGCAACTAGAAAGAATGATCTTCACTTGATTGCCATTTCACATGATT |
| CUST_24974_PI426222305 | JHI_St_60k_v1 | DMT400024087 | CACTTGGAATATCTGCAACTAGAAAGAATGATCTTCACTTGATTGCCATTTCACATGATT |
Microarray Signals from CUST_24842_PI426222305
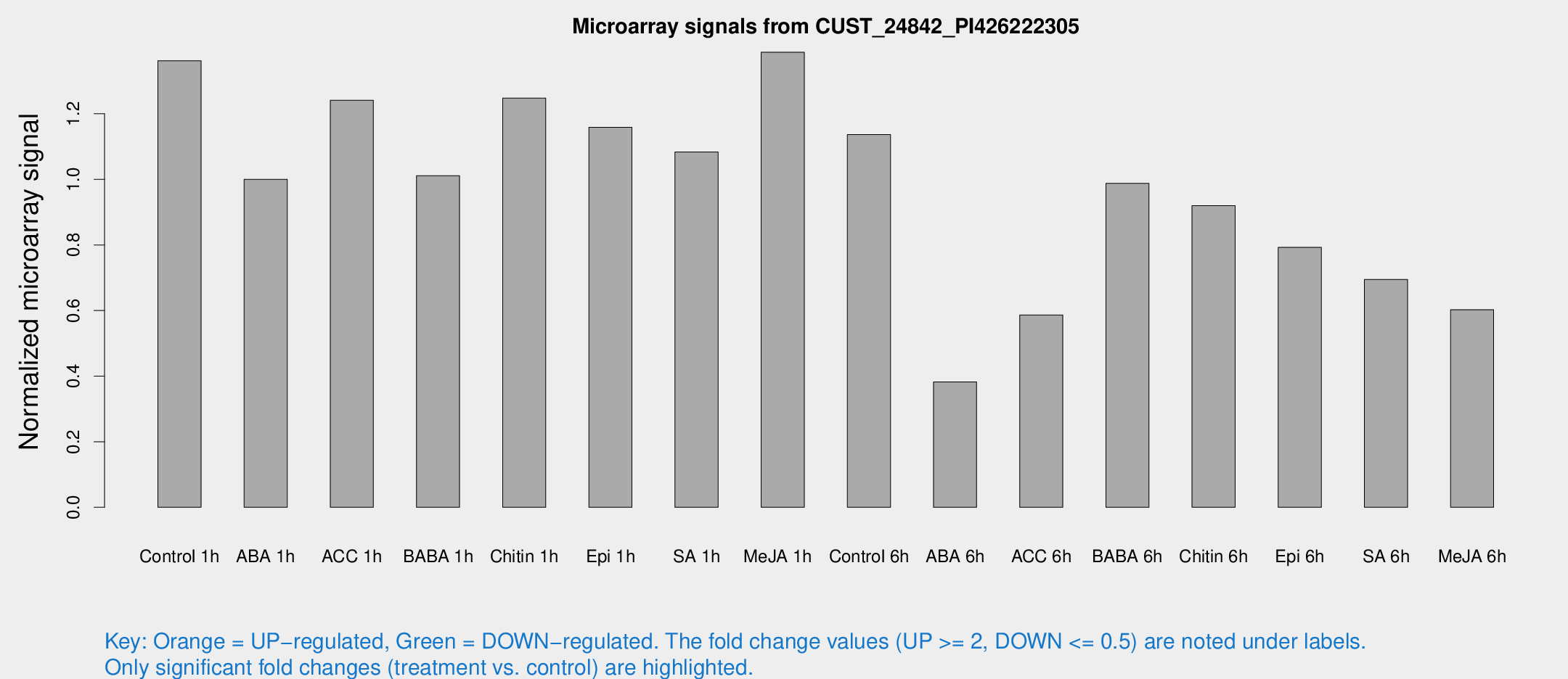
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1055.65 | 130.148 | 1.36162 | 0.0787283 |
| ABA 1h | 697.715 | 130.753 | 1.00007 | 0.126928 |
| ACC 1h | 976.869 | 62.8069 | 1.2409 | 0.0820265 |
| BABA 1h | 769.409 | 132.774 | 1.011 | 0.0887615 |
| Chitin 1h | 875.691 | 130.788 | 1.2476 | 0.169701 |
| Epi 1h | 775.536 | 89.7466 | 1.15878 | 0.0992562 |
| SA 1h | 864.357 | 106.778 | 1.08357 | 0.0935402 |
| Me-JA 1h | 869.693 | 65.916 | 1.38763 | 0.0802814 |
| Control 6h | 1008.65 | 333.093 | 1.13677 | 0.40183 |
| ABA 6h | 310.729 | 18.2868 | 0.382117 | 0.0224501 |
| ACC 6h | 543.432 | 134.727 | 0.58643 | 0.0556021 |
| BABA 6h | 848.439 | 61.4565 | 0.987739 | 0.0691692 |
| Chitin 6h | 754.841 | 73.5138 | 0.919786 | 0.0625902 |
| Epi 6h | 691.562 | 82.252 | 0.792728 | 0.106456 |
| SA 6h | 537.221 | 77.3875 | 0.694927 | 0.0404059 |
| Me-JA 6h | 466.546 | 60.6926 | 0.602399 | 0.0350496 |
Source Transcript PGSC0003DMT400024086 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G26320.1 | +1 | 1e-107 | 330 | 179/454 (39%) | cytochrome P450, family 71, subfamily B, polypeptide 36 | chr3:9644383-9646064 REVERSE LENGTH=500 |
