Probe CUST_24970_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24970_PI426222305 | JHI_St_60k_v1 | DMT400024018 | GAGCATGATTAGGAACTTTCCTGTATCTAATTTTAGGTTTACTTAGGATGTGTATAGAGG |
All Microarray Probes Designed to Gene DMG402009291
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24970_PI426222305 | JHI_St_60k_v1 | DMT400024018 | GAGCATGATTAGGAACTTTCCTGTATCTAATTTTAGGTTTACTTAGGATGTGTATAGAGG |
| CUST_24862_PI426222305 | JHI_St_60k_v1 | DMT400024017 | GAGCATGATTAGGAACTTTCCTGTATCTAATTTTAGGTTTACTTAGGATGTGTATAGAGG |
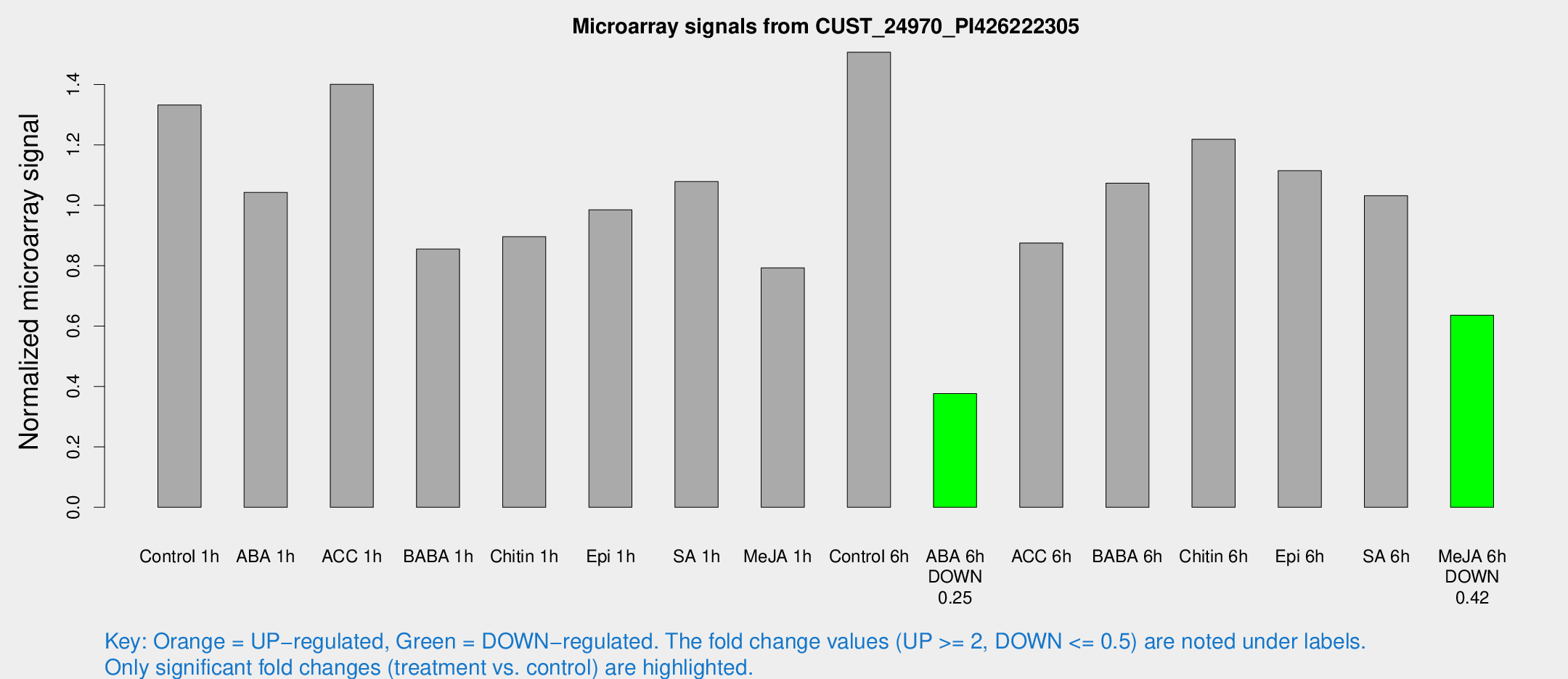
Microarray Signals from CUST_24970_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 13943.4 | 1954.52 | 1.33224 | 0.0980727 |
| ABA 1h | 9536.62 | 803.99 | 1.04262 | 0.126419 |
| ACC 1h | 14784.4 | 854.836 | 1.40073 | 0.0994643 |
| BABA 1h | 8861.7 | 1749.84 | 0.855101 | 0.103076 |
| Chitin 1h | 8462.21 | 1291.6 | 0.896349 | 0.127644 |
| Epi 1h | 8760.06 | 506.352 | 0.985075 | 0.0568744 |
| SA 1h | 11466.7 | 1010.66 | 1.07864 | 0.0622759 |
| Me-JA 1h | 6715.29 | 666.7 | 0.792996 | 0.0457853 |
| Control 6h | 16369.7 | 3541.09 | 1.50677 | 0.224553 |
| ABA 6h | 4142.93 | 372.952 | 0.376655 | 0.0271734 |
| ACC 6h | 10472.6 | 1365.78 | 0.874843 | 0.0505102 |
| BABA 6h | 12611.7 | 1890.2 | 1.07306 | 0.185199 |
| Chitin 6h | 13348 | 772.851 | 1.21827 | 0.070338 |
| Epi 6h | 13060.6 | 1473.77 | 1.11459 | 0.174015 |
| SA 6h | 10691.9 | 1450.53 | 1.03167 | 0.0595643 |
| Me-JA 6h | 6977.08 | 1700.01 | 0.63588 | 0.126894 |
Source Transcript PGSC0003DMT400024018 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G32080.1 | +3 | 0.0 | 686 | 366/476 (77%) | membrane protein, putative | chr1:11537572-11539756 REVERSE LENGTH=512 |
