Probe CUST_24862_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24862_PI426222305 | JHI_St_60k_v1 | DMT400024017 | GAGCATGATTAGGAACTTTCCTGTATCTAATTTTAGGTTTACTTAGGATGTGTATAGAGG |
All Microarray Probes Designed to Gene DMG402009291
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24970_PI426222305 | JHI_St_60k_v1 | DMT400024018 | GAGCATGATTAGGAACTTTCCTGTATCTAATTTTAGGTTTACTTAGGATGTGTATAGAGG |
| CUST_24862_PI426222305 | JHI_St_60k_v1 | DMT400024017 | GAGCATGATTAGGAACTTTCCTGTATCTAATTTTAGGTTTACTTAGGATGTGTATAGAGG |
Microarray Signals from CUST_24862_PI426222305
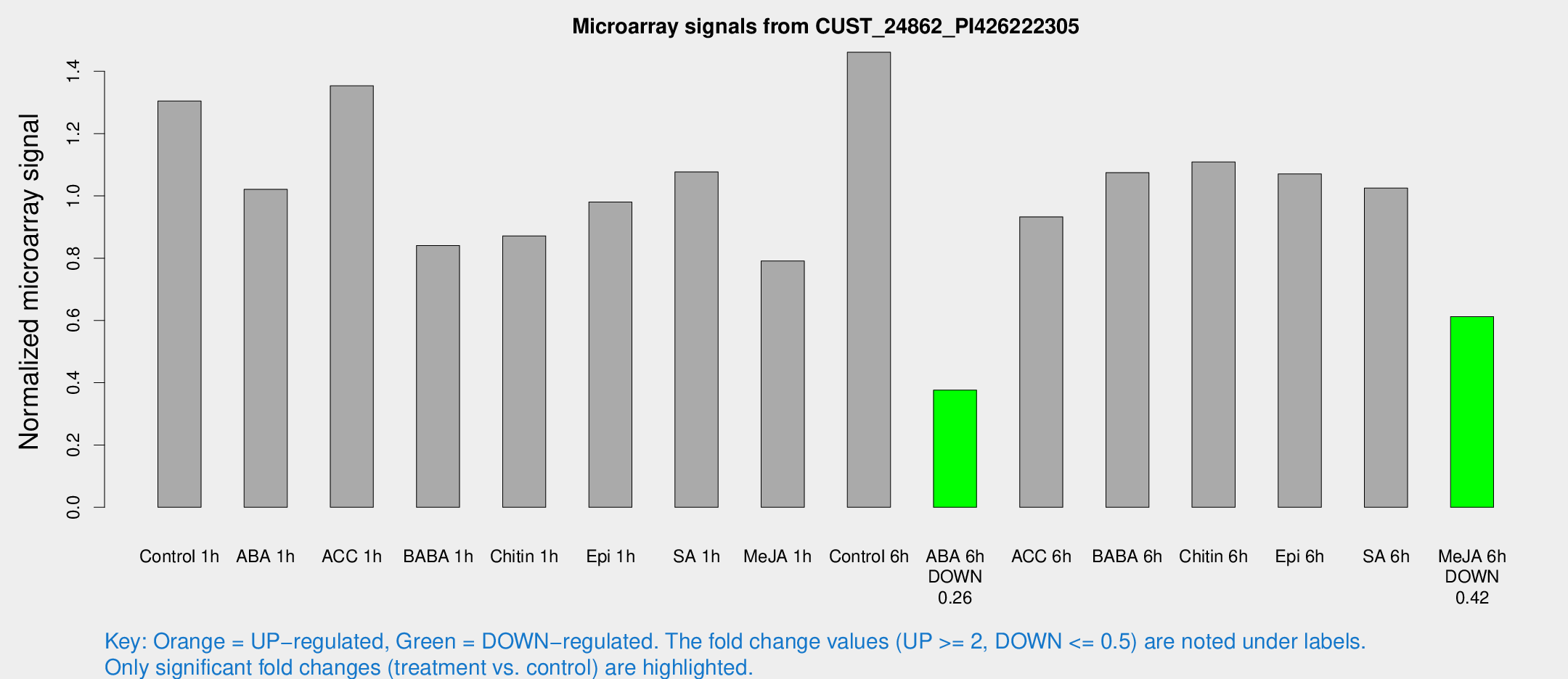
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 13551.7 | 1702.5 | 1.30476 | 0.0782946 |
| ABA 1h | 9309.66 | 816.222 | 1.02111 | 0.121866 |
| ACC 1h | 14213.6 | 820.877 | 1.35353 | 0.0911141 |
| BABA 1h | 8665.4 | 1671.73 | 0.840832 | 0.0972095 |
| Chitin 1h | 8183.9 | 1212.95 | 0.871621 | 0.123067 |
| Epi 1h | 8747.69 | 830.539 | 0.98026 | 0.0654898 |
| SA 1h | 11400.1 | 989.005 | 1.07691 | 0.0621759 |
| Me-JA 1h | 6651.86 | 553.454 | 0.791287 | 0.0456866 |
| Control 6h | 15664.1 | 3150.65 | 1.4614 | 0.183168 |
| ABA 6h | 4121.31 | 369.899 | 0.376314 | 0.0217289 |
| ACC 6h | 11168.3 | 1591.51 | 0.9327 | 0.0567757 |
| BABA 6h | 12659.2 | 2173.51 | 1.07446 | 0.203248 |
| Chitin 6h | 12158 | 1047.15 | 1.1089 | 0.0640231 |
| Epi 6h | 12431.1 | 1039.74 | 1.07061 | 0.161481 |
| SA 6h | 10623.3 | 1576.76 | 1.02519 | 0.0591905 |
| Me-JA 6h | 6695.6 | 1623.3 | 0.612489 | 0.123466 |
Source Transcript PGSC0003DMT400024017 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G32080.1 | +2 | 0.0 | 659 | 353/452 (78%) | membrane protein, putative | chr1:11537572-11539756 REVERSE LENGTH=512 |
