Probe CUST_24630_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24630_PI426222305 | JHI_St_60k_v1 | DMT400054442 | AGCCTAAAAGAAGCTAATAGACCGACACTTGCTTGGTACGCGAAGGAGGATGATAATAGG |
All Microarray Probes Designed to Gene DMG400021131
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24630_PI426222305 | JHI_St_60k_v1 | DMT400054442 | AGCCTAAAAGAAGCTAATAGACCGACACTTGCTTGGTACGCGAAGGAGGATGATAATAGG |
| CUST_24527_PI426222305 | JHI_St_60k_v1 | DMT400054443 | GGGTTAGTAAAGTTTGATCATGGAAATATTTGATTAATTAGGTGTTGCCCATAGAACCAG |
Microarray Signals from CUST_24630_PI426222305
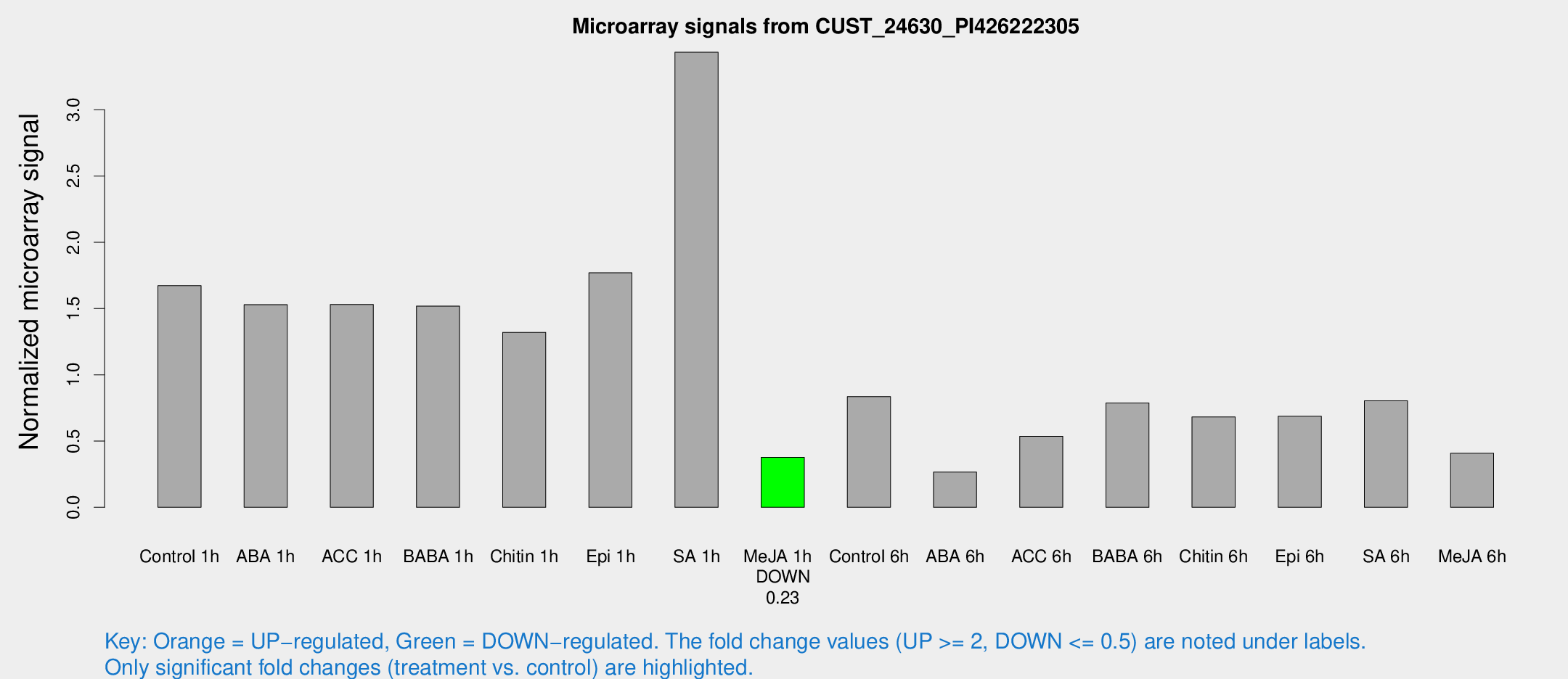
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 677.332 | 92.4091 | 1.67271 | 0.122958 |
| ABA 1h | 542.618 | 52.2088 | 1.52945 | 0.125783 |
| ACC 1h | 649.12 | 124.71 | 1.53069 | 0.215192 |
| BABA 1h | 662.922 | 201.531 | 1.51883 | 0.461928 |
| Chitin 1h | 475.361 | 41.0338 | 1.32035 | 0.0833084 |
| Epi 1h | 609.687 | 35.447 | 1.76955 | 0.102685 |
| SA 1h | 1509.83 | 401.836 | 3.43428 | 0.760402 |
| Me-JA 1h | 123.211 | 11.4706 | 0.376587 | 0.0357515 |
| Control 6h | 380.17 | 132.437 | 0.83545 | 0.270344 |
| ABA 6h | 127.092 | 37.7299 | 0.265859 | 0.0984294 |
| ACC 6h | 254.604 | 47.8547 | 0.535978 | 0.079039 |
| BABA 6h | 372.247 | 86.1104 | 0.788013 | 0.182021 |
| Chitin 6h | 309.224 | 74.8012 | 0.682548 | 0.190672 |
| Epi 6h | 349.639 | 125.895 | 0.687313 | 0.24202 |
| SA 6h | 317.717 | 24.8996 | 0.803479 | 0.126763 |
| Me-JA 6h | 182.075 | 56.4118 | 0.408302 | 0.121562 |
Source Transcript PGSC0003DMT400054442 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G40010.1 | +1 | 8e-42 | 147 | 64/143 (45%) | AAA-ATPase 1 | chr5:16020218-16021762 REVERSE LENGTH=514 |
