Probe CUST_24527_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24527_PI426222305 | JHI_St_60k_v1 | DMT400054443 | GGGTTAGTAAAGTTTGATCATGGAAATATTTGATTAATTAGGTGTTGCCCATAGAACCAG |
All Microarray Probes Designed to Gene DMG400021131
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24630_PI426222305 | JHI_St_60k_v1 | DMT400054442 | AGCCTAAAAGAAGCTAATAGACCGACACTTGCTTGGTACGCGAAGGAGGATGATAATAGG |
| CUST_24527_PI426222305 | JHI_St_60k_v1 | DMT400054443 | GGGTTAGTAAAGTTTGATCATGGAAATATTTGATTAATTAGGTGTTGCCCATAGAACCAG |
Microarray Signals from CUST_24527_PI426222305
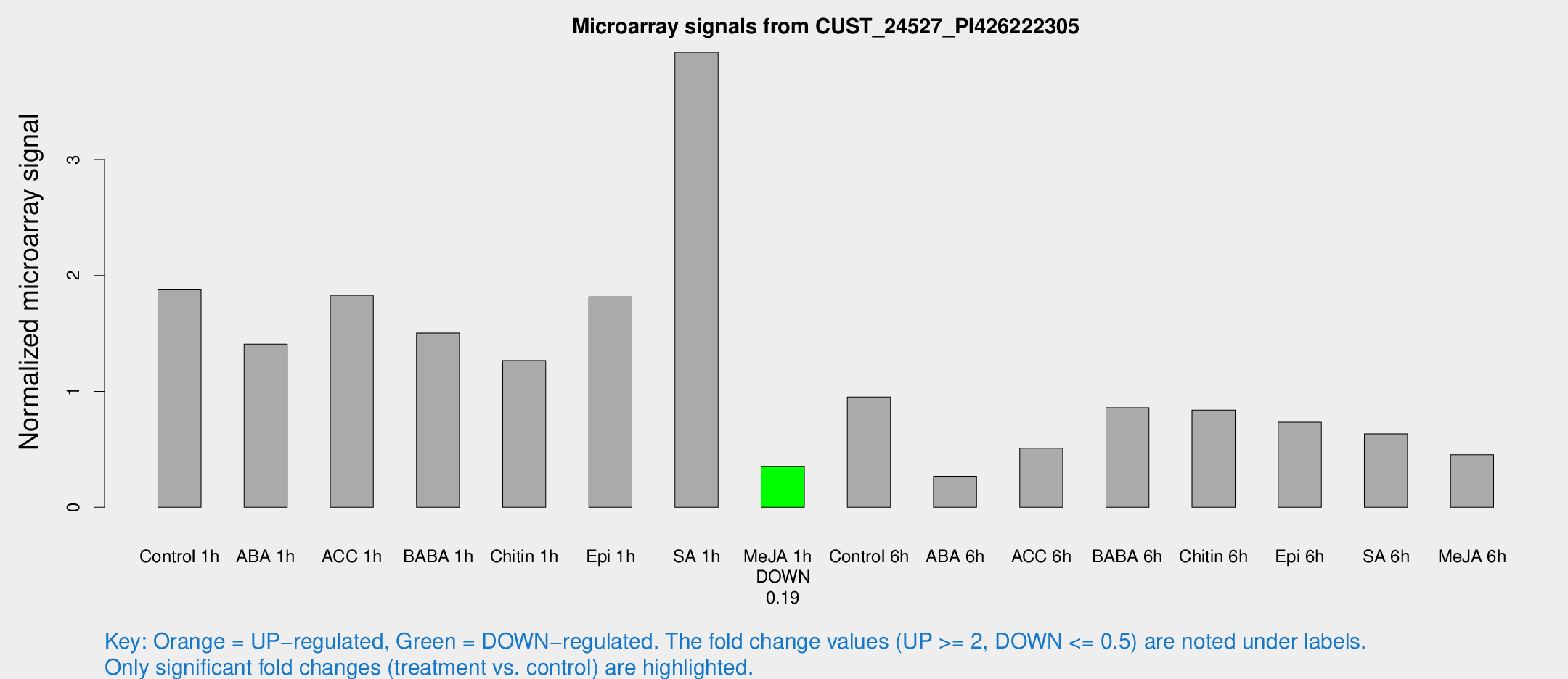
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 4741.54 | 883.211 | 1.87667 | 0.216559 |
| ABA 1h | 3104.81 | 399.147 | 1.40944 | 0.136218 |
| ACC 1h | 4654.35 | 544.878 | 1.8296 | 0.173739 |
| BABA 1h | 4086.18 | 1269.13 | 1.50501 | 0.490989 |
| Chitin 1h | 2818.83 | 317.635 | 1.26667 | 0.0731473 |
| Epi 1h | 3849.58 | 222.616 | 1.81623 | 0.104872 |
| SA 1h | 10638.5 | 2813.99 | 3.92666 | 0.888587 |
| Me-JA 1h | 739.476 | 174.206 | 0.351124 | 0.0511854 |
| Control 6h | 2512.06 | 665.886 | 0.95141 | 0.203543 |
| ABA 6h | 721.226 | 136.327 | 0.268324 | 0.0384163 |
| ACC 6h | 1488.37 | 289.61 | 0.510229 | 0.0471644 |
| BABA 6h | 2418.52 | 379.91 | 0.860115 | 0.118999 |
| Chitin 6h | 2351.54 | 599.924 | 0.839986 | 0.235939 |
| Epi 6h | 2111.41 | 405.898 | 0.734121 | 0.118225 |
| SA 6h | 1902.09 | 692.444 | 0.634468 | 0.285194 |
| Me-JA 6h | 1258.78 | 430.055 | 0.453621 | 0.138496 |
Source Transcript PGSC0003DMT400054443 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G40010.1 | +3 | 0.0 | 536 | 279/467 (60%) | AAA-ATPase 1 | chr5:16020218-16021762 REVERSE LENGTH=514 |
