Probe CUST_24377_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24377_PI426222305 | JHI_St_60k_v1 | DMT400036083 | TAGCTAAGTTAAATCCTTTAAACAGGTAATCACGAATGGGAAGTATAAGAGTGTGATGCA |
All Microarray Probes Designed to Gene DMG400013894
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24334_PI426222305 | JHI_St_60k_v1 | DMT400036081 | GAGAGAGCTGCCACCATGGAAAAGATTAATGATGCTTGTGAAAATTGGGGCTTCTTTGAG |
| CUST_24377_PI426222305 | JHI_St_60k_v1 | DMT400036083 | TAGCTAAGTTAAATCCTTTAAACAGGTAATCACGAATGGGAAGTATAAGAGTGTGATGCA |
Microarray Signals from CUST_24377_PI426222305
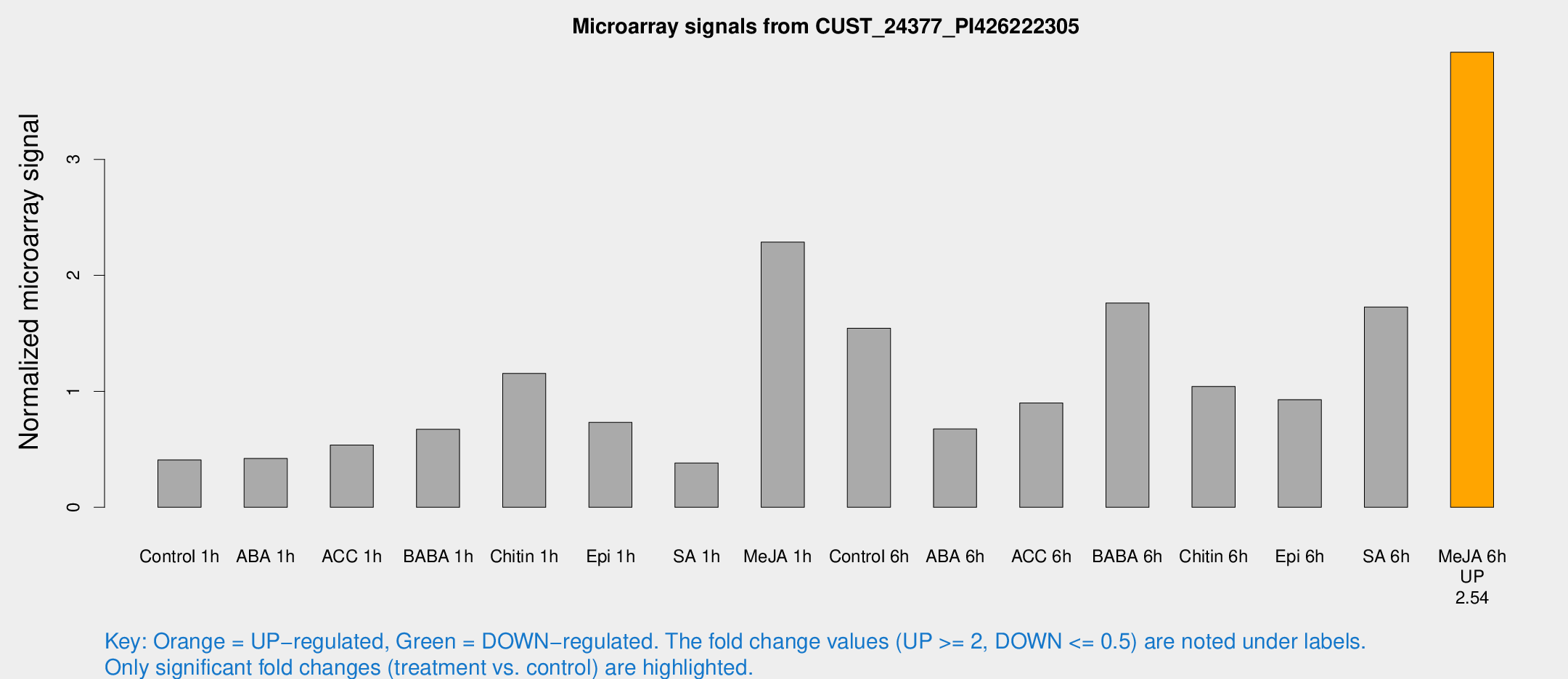
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 5.50263 | 2.96503 | 0.407897 | 0.218689 |
| ABA 1h | 4.93204 | 2.86318 | 0.420382 | 0.237953 |
| ACC 1h | 7.87642 | 3.47063 | 0.537181 | 0.260758 |
| BABA 1h | 11.1864 | 5.67125 | 0.672707 | 0.323311 |
| Chitin 1h | 14.9664 | 3.58038 | 1.15419 | 0.267205 |
| Epi 1h | 8.78433 | 2.96825 | 0.733055 | 0.260528 |
| SA 1h | 5.33428 | 3.0041 | 0.382058 | 0.215284 |
| Me-JA 1h | 27.9994 | 9.10098 | 2.28717 | 0.532216 |
| Control 6h | 21.5076 | 3.28527 | 1.54359 | 0.249051 |
| ABA 6h | 10.9812 | 3.53293 | 0.675904 | 0.255312 |
| ACC 6h | 15.5126 | 5.04939 | 0.899333 | 0.248133 |
| BABA 6h | 27.2389 | 3.77201 | 1.76165 | 0.251433 |
| Chitin 6h | 15.0817 | 3.49044 | 1.04242 | 0.241293 |
| Epi 6h | 15.0661 | 3.67287 | 0.927784 | 0.249936 |
| SA 6h | 23.2269 | 3.49856 | 1.72736 | 0.259983 |
| Me-JA 6h | 53.1113 | 4.3265 | 3.92361 | 0.31938 |
Source Transcript PGSC0003DMT400036083 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G05010.1 | +2 | 1e-35 | 131 | 62/90 (69%) | ethylene-forming enzyme | chr1:1431419-1432695 REVERSE LENGTH=323 |
