Probe CUST_24173_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24173_PI426222305 | JHI_St_60k_v1 | DMT400059046 | CATTTTGTGAAAAGATCTATTTGGTGTGATCGCGTCGTTACCGTAGTATTTTCTTCTCAT |
All Microarray Probes Designed to Gene DMG400022933
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24159_PI426222305 | JHI_St_60k_v1 | DMT400059045 | CATTTTGTGAAAAGATCTATTTGGTGTGATCGCGTCGTTACCGTAGTATTTTCTTCTCAT |
| CUST_24173_PI426222305 | JHI_St_60k_v1 | DMT400059046 | CATTTTGTGAAAAGATCTATTTGGTGTGATCGCGTCGTTACCGTAGTATTTTCTTCTCAT |
Microarray Signals from CUST_24173_PI426222305
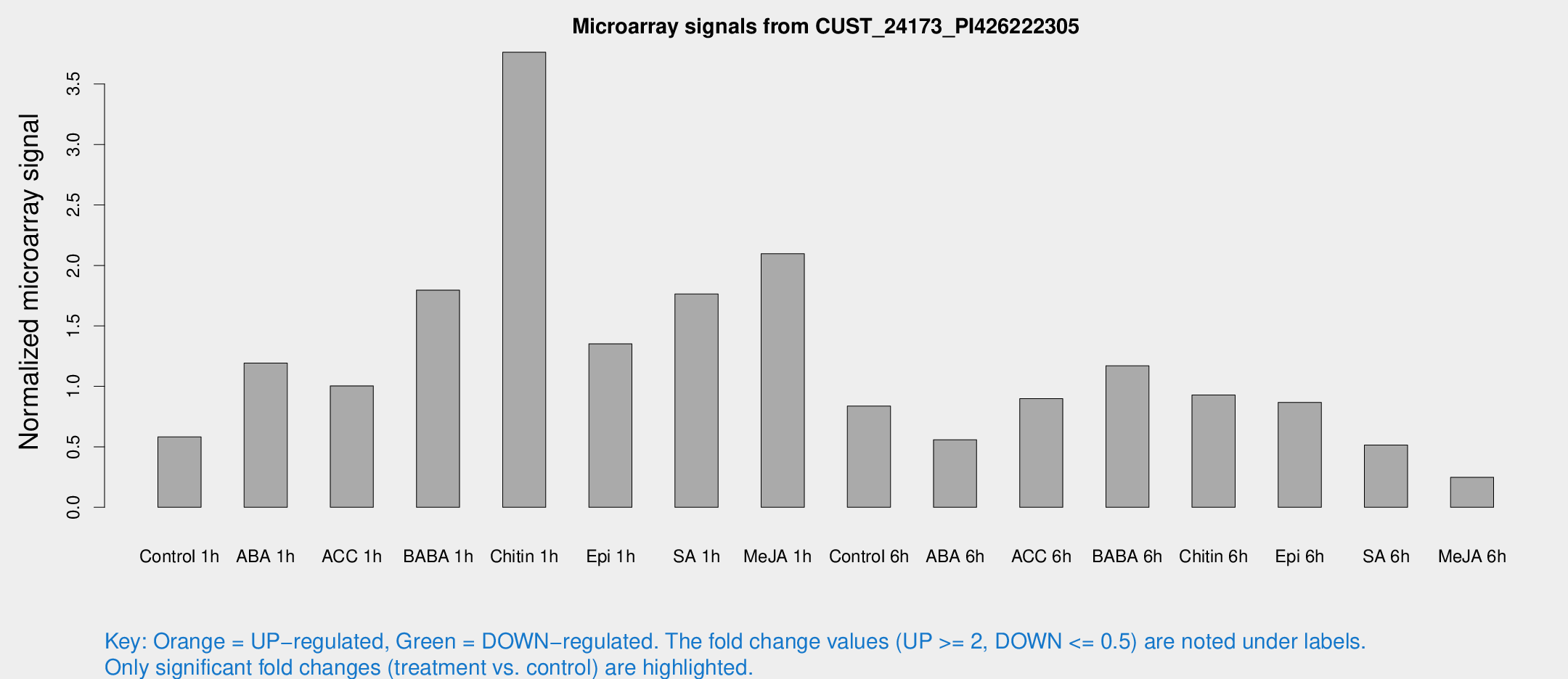
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 58.0007 | 14.0746 | 0.582172 | 0.106399 |
| ABA 1h | 99.1977 | 6.50682 | 1.19223 | 0.0783952 |
| ACC 1h | 97.1643 | 6.59837 | 1.00425 | 0.144027 |
| BABA 1h | 179.928 | 49.9639 | 1.79609 | 0.440934 |
| Chitin 1h | 336.716 | 83.3043 | 3.76273 | 0.882453 |
| Epi 1h | 110.431 | 8.42731 | 1.35143 | 0.113887 |
| SA 1h | 196.493 | 72.7291 | 1.76334 | 0.698997 |
| Me-JA 1h | 164.495 | 24.7093 | 2.09633 | 0.149515 |
| Control 6h | 87.0054 | 27.6264 | 0.836785 | 0.204005 |
| ABA 6h | 60.8697 | 18.9834 | 0.558853 | 0.138896 |
| ACC 6h | 99.5491 | 16.9283 | 0.898375 | 0.0639264 |
| BABA 6h | 124.614 | 13.9713 | 1.17008 | 0.100389 |
| Chitin 6h | 95.4267 | 15.396 | 0.928477 | 0.132988 |
| Epi 6h | 94.467 | 16.665 | 0.867433 | 0.157655 |
| SA 6h | 51.4298 | 13.5686 | 0.515208 | 0.0949439 |
| Me-JA 6h | 33.4635 | 13.9893 | 0.247902 | 0.220954 |
Source Transcript PGSC0003DMT400059046 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G49360.1 | +3 | 0.0 | 1119 | 552/777 (71%) | beta-xylosidase 1 | chr5:20012179-20016659 REVERSE LENGTH=774 |
