Probe CUST_24159_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24159_PI426222305 | JHI_St_60k_v1 | DMT400059045 | CATTTTGTGAAAAGATCTATTTGGTGTGATCGCGTCGTTACCGTAGTATTTTCTTCTCAT |
All Microarray Probes Designed to Gene DMG400022933
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24159_PI426222305 | JHI_St_60k_v1 | DMT400059045 | CATTTTGTGAAAAGATCTATTTGGTGTGATCGCGTCGTTACCGTAGTATTTTCTTCTCAT |
| CUST_24173_PI426222305 | JHI_St_60k_v1 | DMT400059046 | CATTTTGTGAAAAGATCTATTTGGTGTGATCGCGTCGTTACCGTAGTATTTTCTTCTCAT |
Microarray Signals from CUST_24159_PI426222305
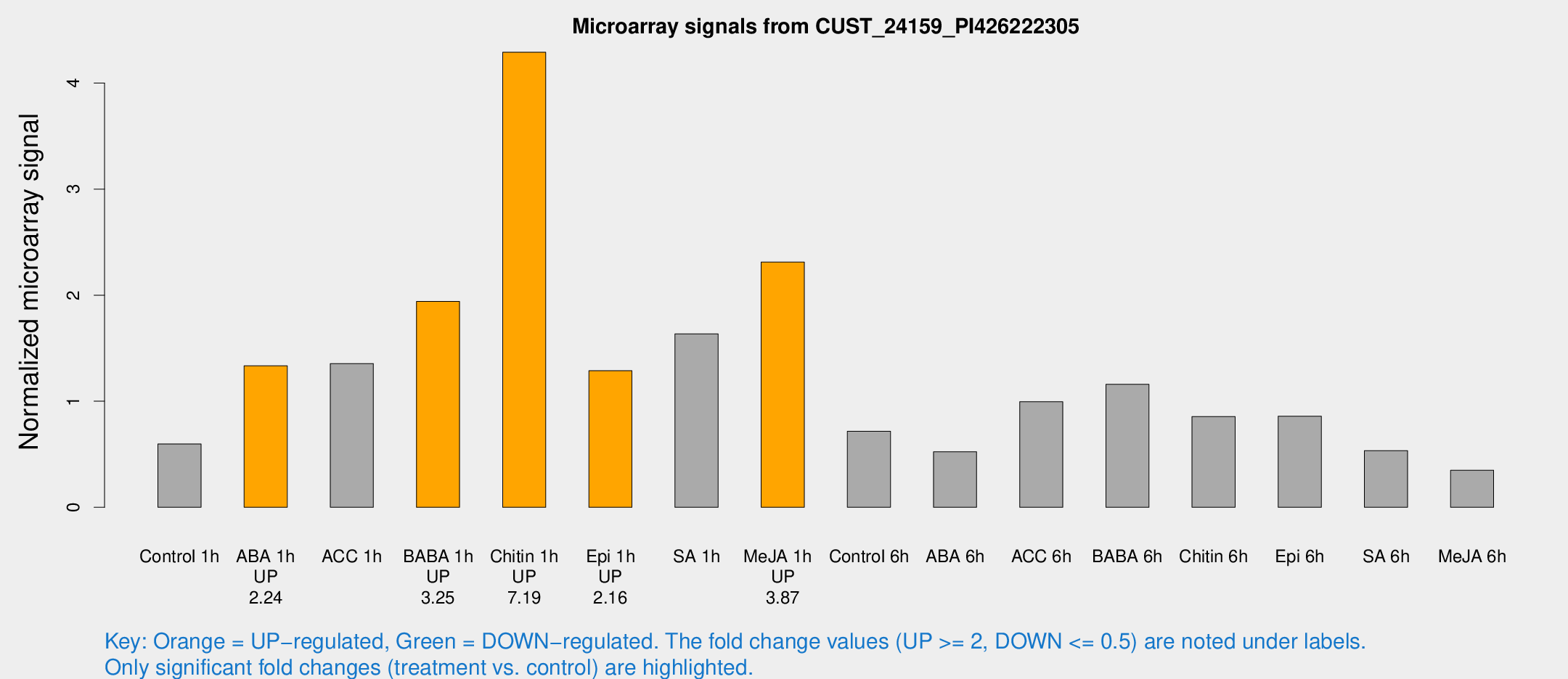
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 53.7346 | 10.75 | 0.596998 | 0.0895738 |
| ABA 1h | 103.47 | 10.9662 | 1.33481 | 0.183643 |
| ACC 1h | 127.167 | 30.5524 | 1.3548 | 0.442029 |
| BABA 1h | 173.902 | 40.5601 | 1.94093 | 0.342424 |
| Chitin 1h | 342.457 | 54.4629 | 4.29059 | 0.783543 |
| Epi 1h | 97.7284 | 10.6072 | 1.28768 | 0.154027 |
| SA 1h | 164.565 | 56.9333 | 1.63574 | 0.56861 |
| Me-JA 1h | 165.163 | 15.4232 | 2.31191 | 0.141799 |
| Control 6h | 74.0837 | 26.3037 | 0.716766 | 0.279345 |
| ABA 6h | 54.5882 | 19.8336 | 0.522942 | 0.162274 |
| ACC 6h | 104.231 | 24.8833 | 0.995072 | 0.0910003 |
| BABA 6h | 115.126 | 16.7233 | 1.15932 | 0.144354 |
| Chitin 6h | 79.2495 | 5.92893 | 0.855901 | 0.0640931 |
| Epi 6h | 90.211 | 24.76 | 0.858112 | 0.270497 |
| SA 6h | 48.4403 | 11.8302 | 0.534095 | 0.0939908 |
| Me-JA 6h | 40.1294 | 16.435 | 0.349504 | 0.227005 |
Source Transcript PGSC0003DMT400059045 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G49360.1 | +3 | 2e-63 | 217 | 112/206 (54%) | beta-xylosidase 1 | chr5:20012179-20016659 REVERSE LENGTH=774 |
