Probe CUST_23381_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23381_PI426222305 | JHI_St_60k_v1 | DMT400002558 | GCTTGTTGTCTGGATTCTGCTGTTGTTGAATTCAAGAGTTTTGTTAACTTGTTTTAGCAT |
All Microarray Probes Designed to Gene DMG400000980
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23381_PI426222305 | JHI_St_60k_v1 | DMT400002558 | GCTTGTTGTCTGGATTCTGCTGTTGTTGAATTCAAGAGTTTTGTTAACTTGTTTTAGCAT |
| CUST_23360_PI426222305 | JHI_St_60k_v1 | DMT400002557 | AAACACTAGAAGATACAATTAGAACTGTTGAAGATGGTCCGGGACAACACCCGACTCAGG |
Microarray Signals from CUST_23381_PI426222305
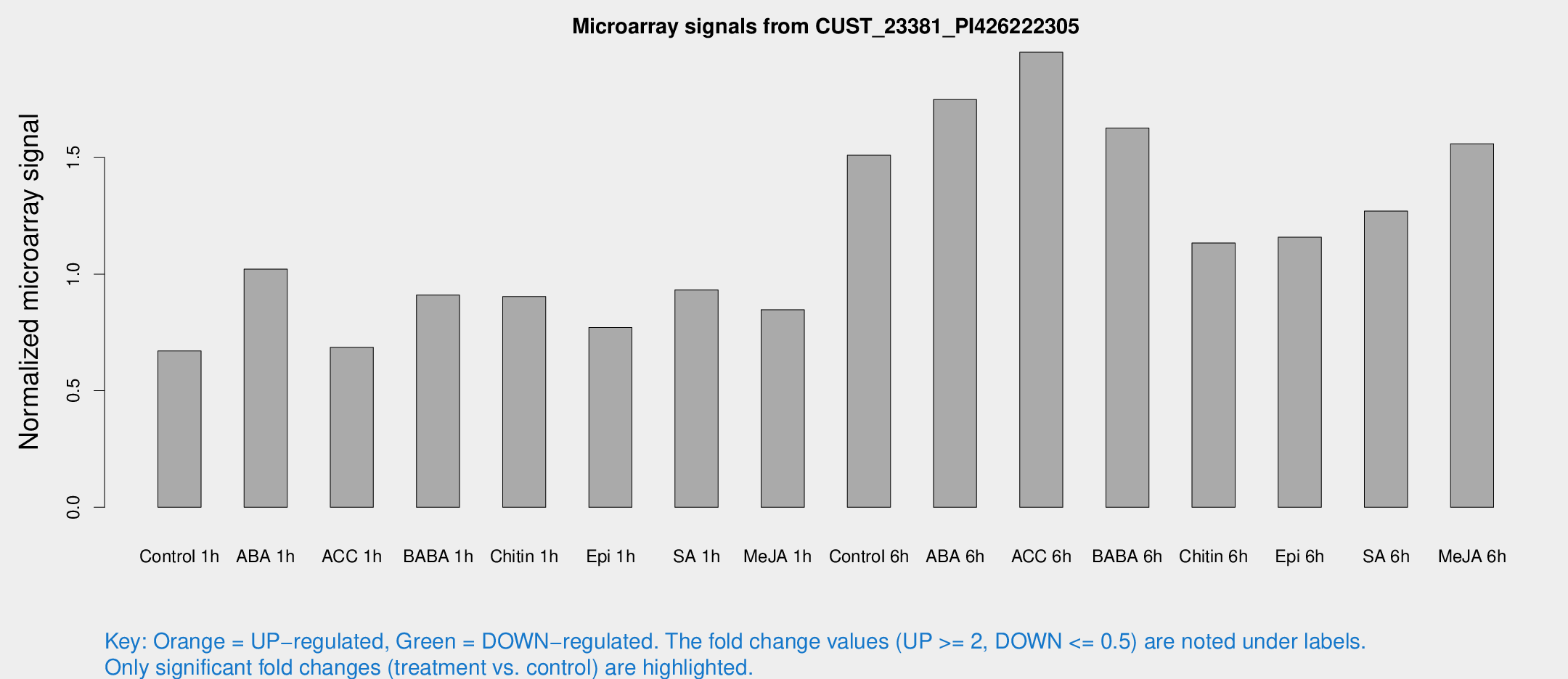
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 557.998 | 57.3196 | 0.670284 | 0.0402035 |
| ABA 1h | 773.939 | 150.957 | 1.02107 | 0.184427 |
| ACC 1h | 595.851 | 93.2699 | 0.686066 | 0.0621479 |
| BABA 1h | 735.592 | 96.9059 | 0.910169 | 0.0527275 |
| Chitin 1h | 676.081 | 67.6601 | 0.90377 | 0.0523462 |
| Epi 1h | 554.875 | 53.5724 | 0.771152 | 0.0685962 |
| SA 1h | 789.947 | 45.8405 | 0.931614 | 0.0539114 |
| Me-JA 1h | 577 | 63.5029 | 0.847548 | 0.0491609 |
| Control 6h | 1357.47 | 357.592 | 1.50973 | 0.347993 |
| ABA 6h | 1533.88 | 88.8514 | 1.7486 | 0.101026 |
| ACC 6h | 1850.72 | 123.696 | 1.95156 | 0.176071 |
| BABA 6h | 1516.37 | 167.657 | 1.62643 | 0.117478 |
| Chitin 6h | 993.944 | 57.537 | 1.13367 | 0.0655751 |
| Epi 6h | 1075.95 | 62.2753 | 1.15858 | 0.0699348 |
| SA 6h | 1091.04 | 227.317 | 1.27054 | 0.157381 |
| Me-JA 6h | 1283.64 | 86.7459 | 1.55908 | 0.090096 |
Source Transcript PGSC0003DMT400002558 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G20330.1 | +1 | 7e-176 | 504 | 280/388 (72%) | PYRIMIDINE B | chr3:7090354-7091904 REVERSE LENGTH=390 |
