Probe CUST_23360_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23360_PI426222305 | JHI_St_60k_v1 | DMT400002557 | AAACACTAGAAGATACAATTAGAACTGTTGAAGATGGTCCGGGACAACACCCGACTCAGG |
All Microarray Probes Designed to Gene DMG400000980
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23381_PI426222305 | JHI_St_60k_v1 | DMT400002558 | GCTTGTTGTCTGGATTCTGCTGTTGTTGAATTCAAGAGTTTTGTTAACTTGTTTTAGCAT |
| CUST_23360_PI426222305 | JHI_St_60k_v1 | DMT400002557 | AAACACTAGAAGATACAATTAGAACTGTTGAAGATGGTCCGGGACAACACCCGACTCAGG |
Microarray Signals from CUST_23360_PI426222305
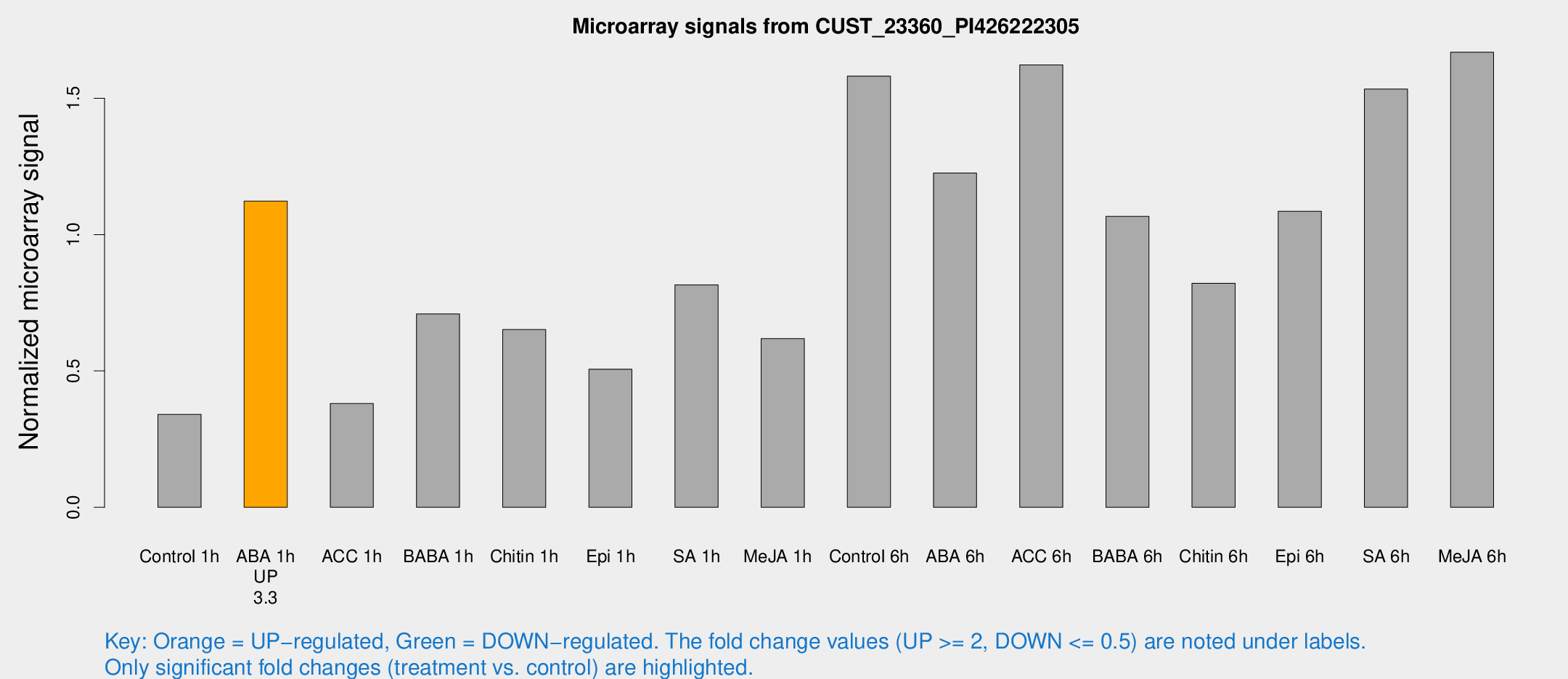
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 12.5244 | 4.19676 | 0.340627 | 0.122961 |
| ABA 1h | 33.1817 | 4.13889 | 1.1227 | 0.143876 |
| ACC 1h | 15.8618 | 7.21322 | 0.380537 | 0.182735 |
| BABA 1h | 22.863 | 4.17734 | 0.709323 | 0.133398 |
| Chitin 1h | 19.8533 | 3.67043 | 0.651891 | 0.123748 |
| Epi 1h | 14.6314 | 3.70113 | 0.506472 | 0.131428 |
| SA 1h | 28.7546 | 5.10972 | 0.815774 | 0.185567 |
| Me-JA 1h | 17.5357 | 3.98482 | 0.618688 | 0.146941 |
| Control 6h | 59.3285 | 17.9767 | 1.58107 | 0.470645 |
| ABA 6h | 44.585 | 7.92262 | 1.2256 | 0.14367 |
| ACC 6h | 62.1799 | 5.78321 | 1.62255 | 0.148085 |
| BABA 6h | 40.0228 | 4.96431 | 1.06693 | 0.132189 |
| Chitin 6h | 30.9616 | 8.05136 | 0.821832 | 0.190598 |
| Epi 6h | 41.0354 | 5.25072 | 1.08581 | 0.151699 |
| SA 6h | 51.5164 | 7.53727 | 1.53378 | 0.297437 |
| Me-JA 6h | 56.1542 | 7.02634 | 1.66879 | 0.151131 |
Source Transcript PGSC0003DMT400002557 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G20330.1 | +1 | 6e-157 | 449 | 241/333 (72%) | PYRIMIDINE B | chr3:7090354-7091904 REVERSE LENGTH=390 |
