Probe CUST_23212_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23212_PI426222305 | JHI_St_60k_v1 | DMT400026722 | AAATTTATGTTGTTGAGGTCTCATGTTGGAGCAGAGATTGCACTGTTGAGTTTTCTTGCT |
All Microarray Probes Designed to Gene DMG400010326
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23176_PI426222305 | JHI_St_60k_v1 | DMT400026727 | TTGTTCTCTGTGTCGACTCTTTCCAGAGCATGTACACAAATGCTTTGTGGTCTACTTCGA |
| CUST_23212_PI426222305 | JHI_St_60k_v1 | DMT400026722 | AAATTTATGTTGTTGAGGTCTCATGTTGGAGCAGAGATTGCACTGTTGAGTTTTCTTGCT |
Microarray Signals from CUST_23212_PI426222305
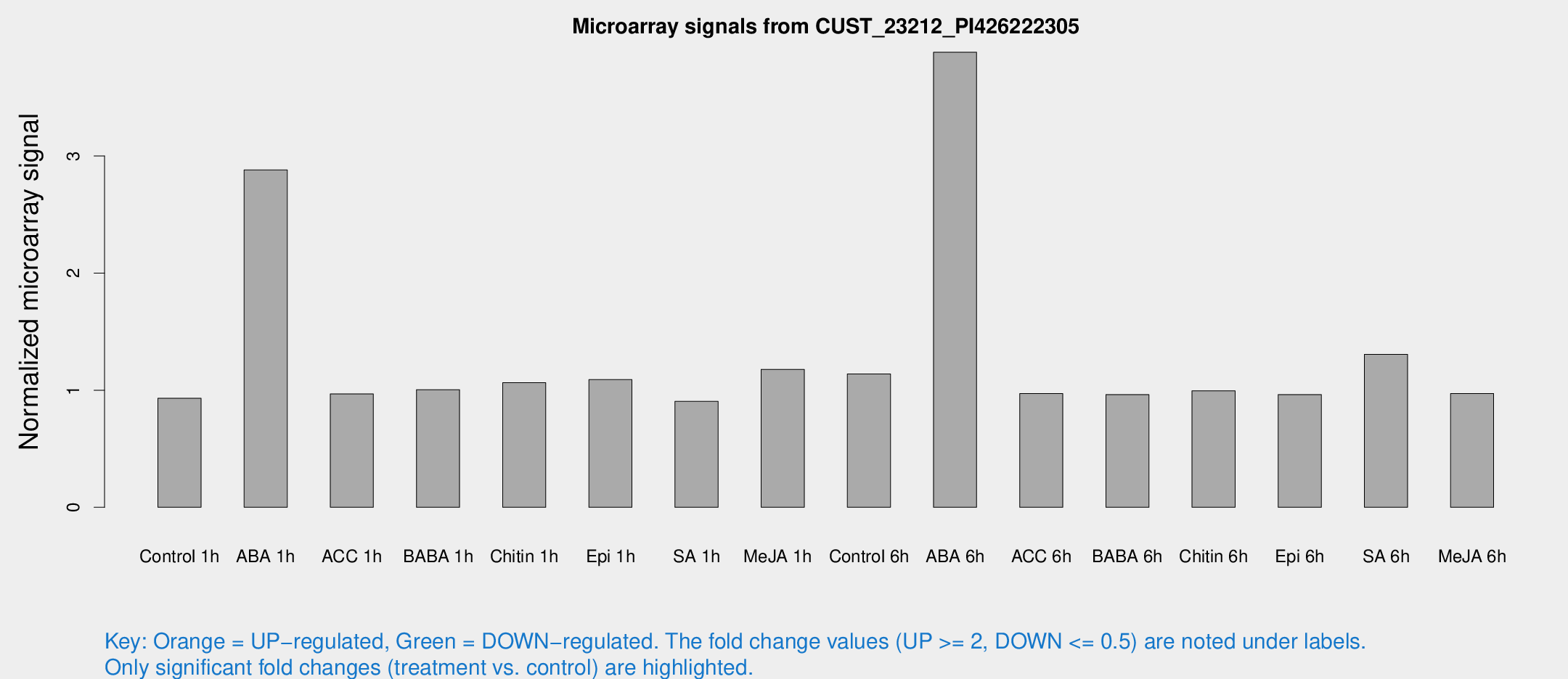
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 4.93325 | 2.85816 | 0.931048 | 0.528506 |
| ABA 1h | 16.8718 | 6.06329 | 2.88085 | 1.51152 |
| ACC 1h | 5.37751 | 3.11747 | 0.967827 | 0.560443 |
| BABA 1h | 5.23943 | 3.03893 | 1.00478 | 0.582211 |
| Chitin 1h | 4.98064 | 2.89663 | 1.06461 | 0.592017 |
| Epi 1h | 4.96484 | 2.88007 | 1.09057 | 0.614039 |
| SA 1h | 4.98932 | 2.88943 | 0.90461 | 0.520391 |
| Me-JA 1h | 5.08759 | 2.95413 | 1.17833 | 0.665983 |
| Control 6h | 6.39621 | 3.01436 | 1.13828 | 0.574277 |
| ABA 6h | 22.6635 | 3.39444 | 3.88664 | 0.64338 |
| ACC 6h | 6.14895 | 3.64102 | 0.970986 | 0.562269 |
| BABA 6h | 5.83589 | 3.38727 | 0.962523 | 0.557402 |
| Chitin 6h | 5.72749 | 3.31693 | 0.995397 | 0.576359 |
| Epi 6h | 5.9142 | 3.45183 | 0.96307 | 0.557672 |
| SA 6h | 7.34433 | 3.19776 | 1.30613 | 0.635946 |
| Me-JA 6h | 5.19749 | 3.0122 | 0.971522 | 0.558768 |
Source Transcript PGSC0003DMT400026722 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G14960.1 | +1 | 3e-10 | 59 | 25/31 (81%) | Auxin-responsive GH3 family protein | chr2:6451659-6453670 REVERSE LENGTH=590 |
