Probe CUST_23176_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23176_PI426222305 | JHI_St_60k_v1 | DMT400026727 | TTGTTCTCTGTGTCGACTCTTTCCAGAGCATGTACACAAATGCTTTGTGGTCTACTTCGA |
All Microarray Probes Designed to Gene DMG400010326
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23176_PI426222305 | JHI_St_60k_v1 | DMT400026727 | TTGTTCTCTGTGTCGACTCTTTCCAGAGCATGTACACAAATGCTTTGTGGTCTACTTCGA |
| CUST_23212_PI426222305 | JHI_St_60k_v1 | DMT400026722 | AAATTTATGTTGTTGAGGTCTCATGTTGGAGCAGAGATTGCACTGTTGAGTTTTCTTGCT |
Microarray Signals from CUST_23176_PI426222305
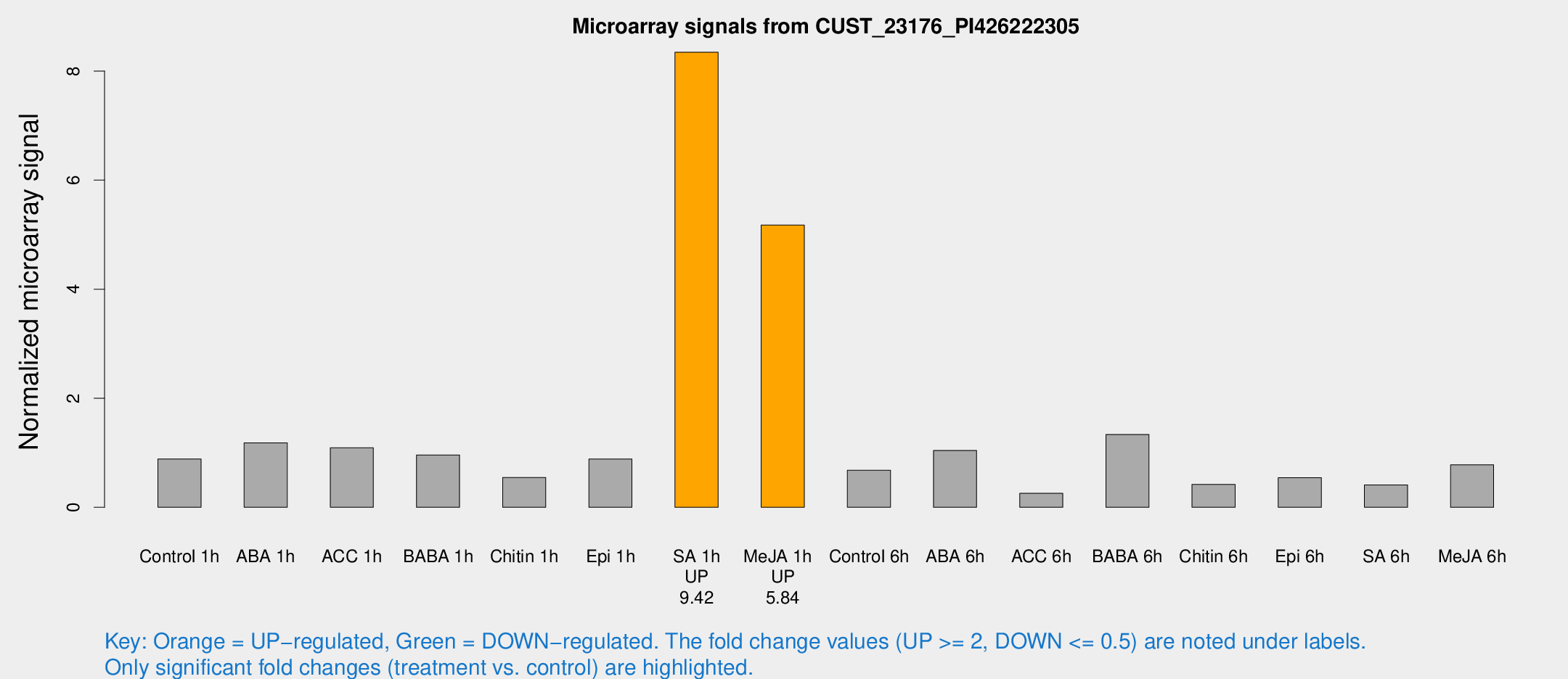
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 26.8142 | 10.8966 | 0.885955 | 0.313723 |
| ABA 1h | 29.486 | 7.90755 | 1.18014 | 0.281494 |
| ACC 1h | 35.3414 | 12.1978 | 1.09084 | 0.510056 |
| BABA 1h | 30.3141 | 13.6596 | 0.959021 | 0.443672 |
| Chitin 1h | 13.9334 | 4.19835 | 0.547777 | 0.18292 |
| Epi 1h | 20.0964 | 3.36651 | 0.884543 | 0.152422 |
| SA 1h | 237.193 | 52.9166 | 8.34529 | 1.76424 |
| Me-JA 1h | 124.646 | 42.6905 | 5.17576 | 1.50205 |
| Control 6h | 24.1884 | 12.8404 | 0.677927 | 0.399017 |
| ABA 6h | 30.4452 | 6.51026 | 1.04263 | 0.206575 |
| ACC 6h | 7.70769 | 4.08301 | 0.25647 | 0.131694 |
| BABA 6h | 42.4694 | 13.2703 | 1.33567 | 0.360517 |
| Chitin 6h | 14.9373 | 7.77306 | 0.418889 | 0.264936 |
| Epi 6h | 20.0158 | 9.5557 | 0.543657 | 0.32752 |
| SA 6h | 13.8372 | 7.44286 | 0.409609 | 0.35113 |
| Me-JA 6h | 27.6593 | 11.0895 | 0.780025 | 0.584755 |
Source Transcript PGSC0003DMT400026727 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G14960.1 | +1 | 7e-49 | 66 | 38/71 (54%) | Auxin-responsive GH3 family protein | chr2:6451659-6453670 REVERSE LENGTH=590 |
