Probe CUST_23107_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23107_PI426222305 | JHI_St_60k_v1 | DMT400021052 | GCTAAGGCTGTGTTGCAACAAAAAGTGGTCAAATCTCGAAAGATGGCACCCTCCTCAATT |
All Microarray Probes Designed to Gene DMG400008148
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22982_PI426222305 | JHI_St_60k_v1 | DMT400021053 | GGCGAAGTGGAAAGTCCAATGTTTCAAAAATCGGATTTAATCTTTATATGGTTGAGATAC |
| CUST_23107_PI426222305 | JHI_St_60k_v1 | DMT400021052 | GCTAAGGCTGTGTTGCAACAAAAAGTGGTCAAATCTCGAAAGATGGCACCCTCCTCAATT |
Microarray Signals from CUST_23107_PI426222305
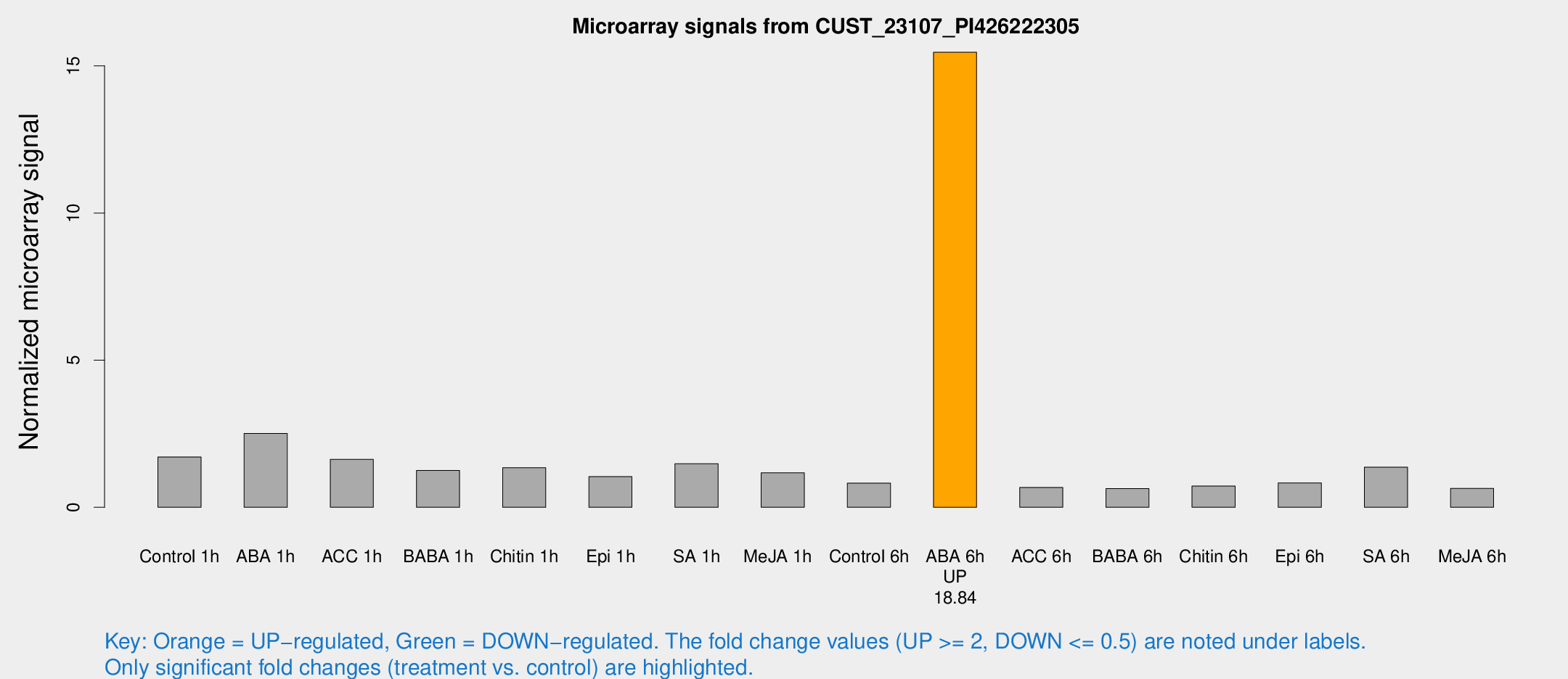
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 18.2763 | 5.16716 | 1.71055 | 0.678384 |
| ABA 1h | 22.6559 | 5.74183 | 2.50848 | 0.584276 |
| ACC 1h | 17.5686 | 5.53722 | 1.63043 | 0.678413 |
| BABA 1h | 12.6337 | 3.73305 | 1.25442 | 0.538564 |
| Chitin 1h | 12.8014 | 3.58688 | 1.34588 | 0.538062 |
| Epi 1h | 8.82636 | 3.38226 | 1.04067 | 0.429372 |
| SA 1h | 15.1486 | 3.60451 | 1.48005 | 0.407093 |
| Me-JA 1h | 9.81254 | 3.63602 | 1.17171 | 0.492287 |
| Control 6h | 8.07865 | 3.5682 | 0.820768 | 0.385342 |
| ABA 6h | 163.343 | 29.3258 | 15.4649 | 2.46111 |
| ACC 6h | 7.5648 | 4.41084 | 0.674834 | 0.382068 |
| BABA 6h | 6.80505 | 3.9545 | 0.634171 | 0.367623 |
| Chitin 6h | 7.36443 | 3.92876 | 0.720178 | 0.388272 |
| Epi 6h | 9.88987 | 4.30948 | 0.830689 | 0.410447 |
| SA 6h | 14.2297 | 4.03623 | 1.36303 | 0.48741 |
| Me-JA 6h | 6.15507 | 3.57061 | 0.645817 | 0.374423 |
Source Transcript PGSC0003DMT400021052 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G20910.1 | +1 | 1e-12 | 62 | 29/46 (63%) | nuclear factor Y, subunit A9 | chr3:7326495-7328369 FORWARD LENGTH=303 |
