Probe CUST_22982_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22982_PI426222305 | JHI_St_60k_v1 | DMT400021053 | GGCGAAGTGGAAAGTCCAATGTTTCAAAAATCGGATTTAATCTTTATATGGTTGAGATAC |
All Microarray Probes Designed to Gene DMG400008148
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22982_PI426222305 | JHI_St_60k_v1 | DMT400021053 | GGCGAAGTGGAAAGTCCAATGTTTCAAAAATCGGATTTAATCTTTATATGGTTGAGATAC |
| CUST_23107_PI426222305 | JHI_St_60k_v1 | DMT400021052 | GCTAAGGCTGTGTTGCAACAAAAAGTGGTCAAATCTCGAAAGATGGCACCCTCCTCAATT |
Microarray Signals from CUST_22982_PI426222305
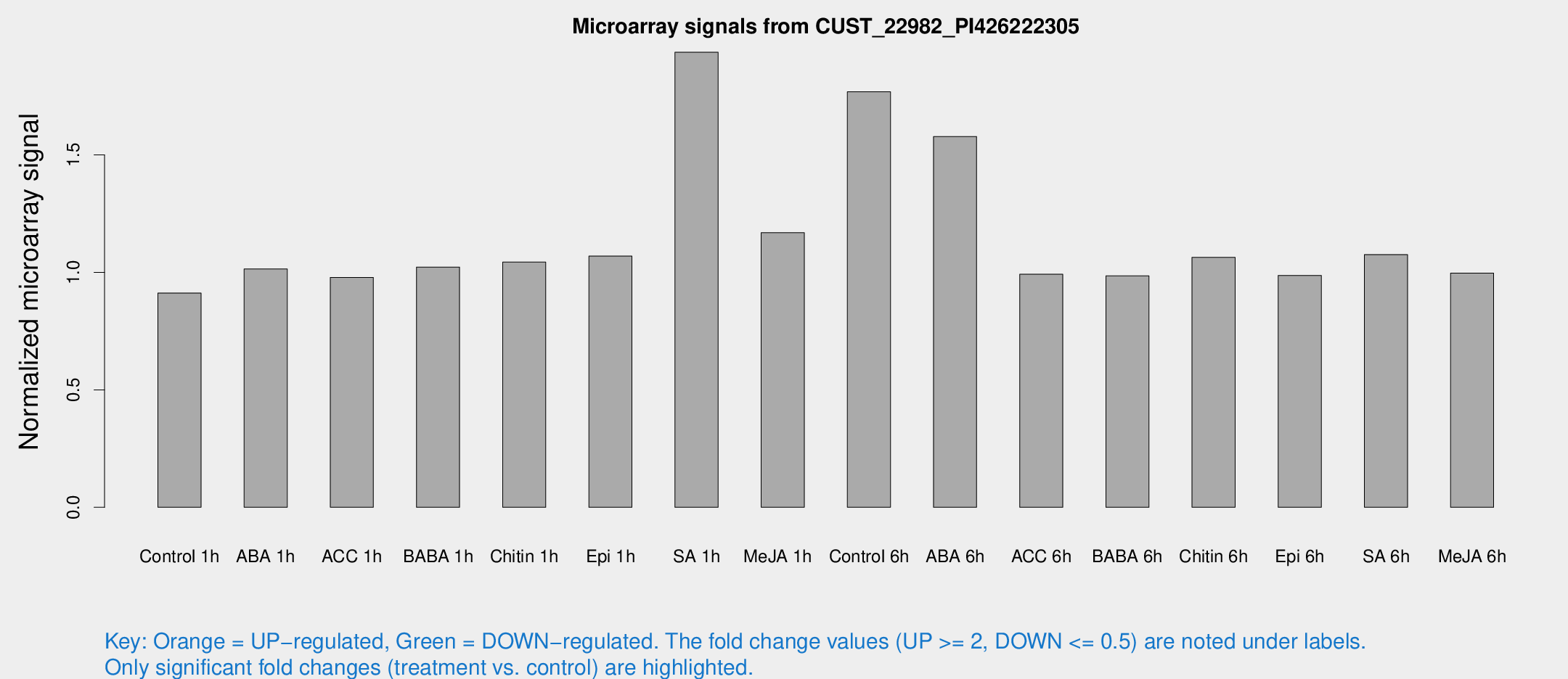
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 5.64614 | 3.27084 | 0.912294 | 0.528297 |
| ABA 1h | 5.56064 | 3.22707 | 1.01466 | 0.588037 |
| ACC 1h | 6.22798 | 3.61247 | 0.978714 | 0.566809 |
| BABA 1h | 6.10231 | 3.53552 | 1.02275 | 0.592229 |
| Chitin 1h | 5.8267 | 3.38621 | 1.04379 | 0.604589 |
| Epi 1h | 5.73562 | 3.32947 | 1.06917 | 0.619433 |
| SA 1h | 22.1427 | 16.3039 | 1.93728 | 2.71273 |
| Me-JA 1h | 5.91622 | 3.43431 | 1.16878 | 0.676912 |
| Control 6h | 15.5733 | 9.37166 | 1.76867 | 1.22551 |
| ABA 6h | 11.1366 | 3.71649 | 1.57837 | 0.6135 |
| ACC 6h | 7.2191 | 4.29081 | 0.992297 | 0.57461 |
| BABA 6h | 6.84247 | 3.97389 | 0.985206 | 0.570514 |
| Chitin 6h | 7.02056 | 3.87001 | 1.06393 | 0.588987 |
| Epi 6h | 6.94402 | 4.0558 | 0.987036 | 0.571549 |
| SA 6h | 6.59912 | 3.72454 | 1.07592 | 0.60841 |
| Me-JA 6h | 6.14779 | 3.56514 | 0.996971 | 0.577612 |
Source Transcript PGSC0003DMT400021053 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G12840.1 | +3 | 1e-23 | 102 | 46/66 (70%) | nuclear factor Y, subunit A1 | chr5:4051147-4052961 REVERSE LENGTH=272 |
