Probe CUST_23072_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23072_PI426222305 | JHI_St_60k_v1 | DMT400060816 | GTCAGCTATCATCATACACAATTACAATATCCAAGTAGTGGAACCAGAAAATGTTTATCC |
All Microarray Probes Designed to Gene DMG400023657
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23072_PI426222305 | JHI_St_60k_v1 | DMT400060816 | GTCAGCTATCATCATACACAATTACAATATCCAAGTAGTGGAACCAGAAAATGTTTATCC |
| CUST_22975_PI426222305 | JHI_St_60k_v1 | DMT400060817 | GTCAGCTATCATCATACACAATTACAATATCCAAGTAGTGGAACCAGAAAATGTTTATCC |
Microarray Signals from CUST_23072_PI426222305
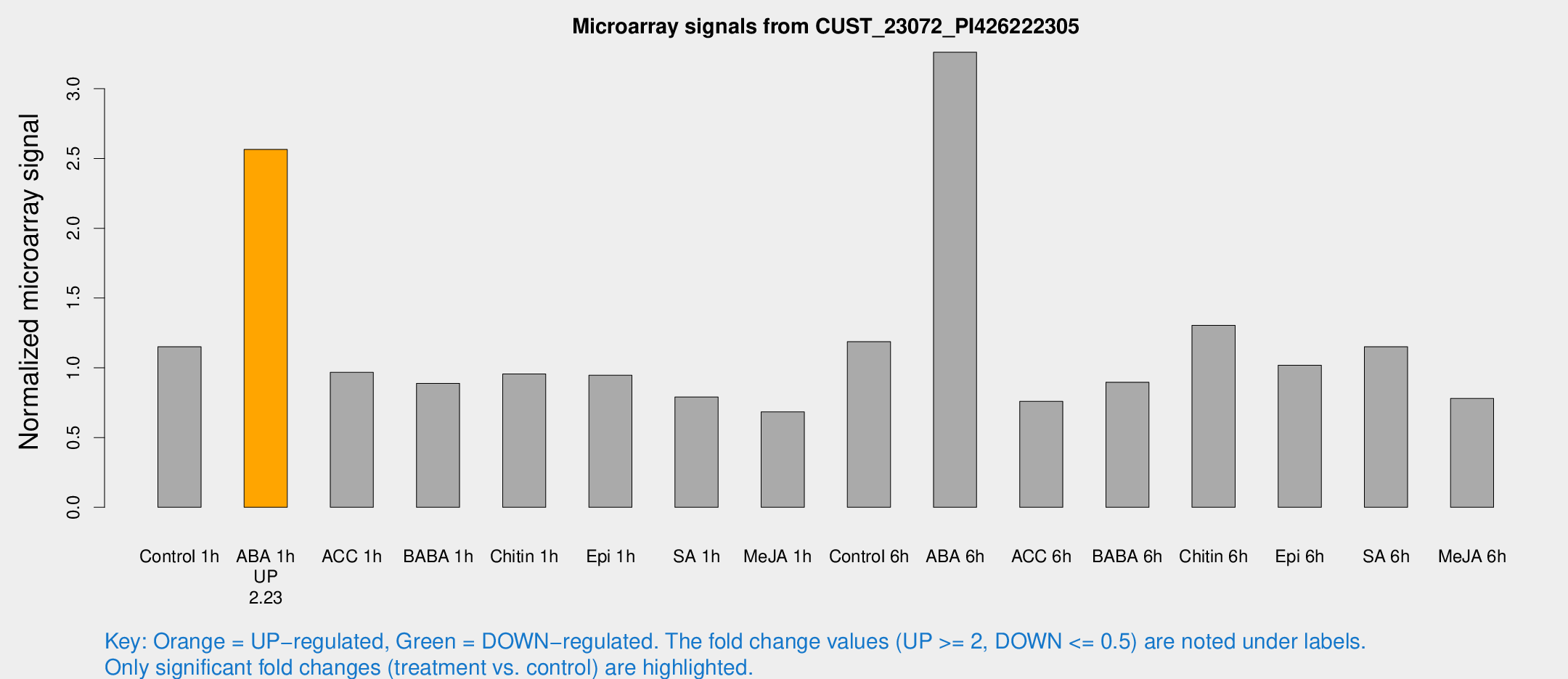
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 703.237 | 40.7806 | 1.15055 | 0.0667088 |
| ABA 1h | 1398.84 | 128.31 | 2.56476 | 0.148226 |
| ACC 1h | 684.126 | 224.6 | 0.966846 | 0.289082 |
| BABA 1h | 523.49 | 30.4903 | 0.888423 | 0.110366 |
| Chitin 1h | 533.447 | 69.7688 | 0.95519 | 0.155543 |
| Epi 1h | 500.766 | 29.15 | 0.946245 | 0.0550503 |
| SA 1h | 499.724 | 39.5739 | 0.791117 | 0.0918457 |
| Me-JA 1h | 344.608 | 35.282 | 0.683528 | 0.132844 |
| Control 6h | 819.919 | 247.404 | 1.18705 | 0.355297 |
| ABA 6h | 2141.53 | 211.494 | 3.26143 | 0.207043 |
| ACC 6h | 539.155 | 59.3686 | 0.759573 | 0.0712356 |
| BABA 6h | 657.572 | 168.624 | 0.896001 | 0.227078 |
| Chitin 6h | 853.725 | 68.4152 | 1.30423 | 0.10423 |
| Epi 6h | 734.297 | 146.025 | 1.01846 | 0.318144 |
| SA 6h | 697.371 | 40.5883 | 1.15038 | 0.191406 |
| Me-JA 6h | 558.69 | 235.69 | 0.779698 | 0.268252 |
Source Transcript PGSC0003DMT400060816 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G23180.1 | +1 | 8e-110 | 335 | 173/384 (45%) | cytochrome P450, family 96, subfamily A, polypeptide 1 | chr2:9874953-9876503 FORWARD LENGTH=516 |
