Probe CUST_22975_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22975_PI426222305 | JHI_St_60k_v1 | DMT400060817 | GTCAGCTATCATCATACACAATTACAATATCCAAGTAGTGGAACCAGAAAATGTTTATCC |
All Microarray Probes Designed to Gene DMG400023657
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23072_PI426222305 | JHI_St_60k_v1 | DMT400060816 | GTCAGCTATCATCATACACAATTACAATATCCAAGTAGTGGAACCAGAAAATGTTTATCC |
| CUST_22975_PI426222305 | JHI_St_60k_v1 | DMT400060817 | GTCAGCTATCATCATACACAATTACAATATCCAAGTAGTGGAACCAGAAAATGTTTATCC |
Microarray Signals from CUST_22975_PI426222305
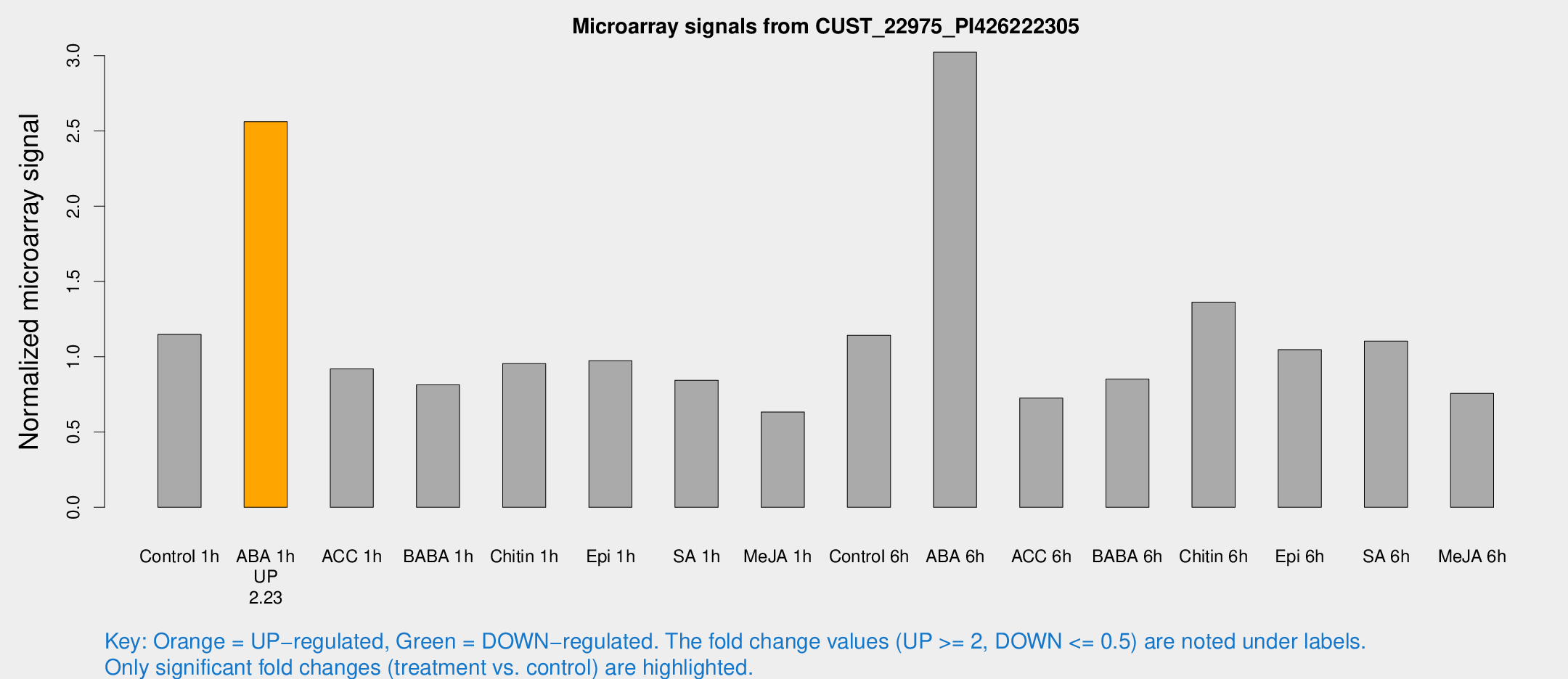
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 717.529 | 41.6892 | 1.14888 | 0.0665585 |
| ABA 1h | 1420.02 | 109.545 | 2.56202 | 0.148046 |
| ACC 1h | 674.605 | 232.328 | 0.919395 | 0.30616 |
| BABA 1h | 489.643 | 28.5596 | 0.814183 | 0.0936826 |
| Chitin 1h | 540.758 | 59.3165 | 0.954745 | 0.111416 |
| Epi 1h | 526.621 | 30.6992 | 0.97395 | 0.0566042 |
| SA 1h | 544.132 | 51.7344 | 0.84337 | 0.101652 |
| Me-JA 1h | 330.302 | 52.7338 | 0.632684 | 0.167373 |
| Control 6h | 796.3 | 227.104 | 1.14195 | 0.32019 |
| ABA 6h | 2015.82 | 170.049 | 3.02317 | 0.174634 |
| ACC 6h | 526.516 | 67.0303 | 0.725635 | 0.0474658 |
| BABA 6h | 637.158 | 175.614 | 0.851943 | 0.218503 |
| Chitin 6h | 911.303 | 86.8938 | 1.36309 | 0.13635 |
| Epi 6h | 750.775 | 107.326 | 1.04772 | 0.238989 |
| SA 6h | 680.503 | 39.5116 | 1.10358 | 0.161573 |
| Me-JA 6h | 576.891 | 272.303 | 0.756771 | 0.307819 |
Source Transcript PGSC0003DMT400060817 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G23180.1 | +3 | 2e-155 | 458 | 239/513 (47%) | cytochrome P450, family 96, subfamily A, polypeptide 1 | chr2:9874953-9876503 FORWARD LENGTH=516 |
