Probe CUST_23008_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23008_PI426222305 | JHI_St_60k_v1 | DMT400060843 | TAGGCCTGACAGTGGTTACTTGAAGATGAAATGGAAACCCAATGCATGTAGTCTTCCTAG |
All Microarray Probes Designed to Gene DMG400023668
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23008_PI426222305 | JHI_St_60k_v1 | DMT400060843 | TAGGCCTGACAGTGGTTACTTGAAGATGAAATGGAAACCCAATGCATGTAGTCTTCCTAG |
| CUST_22955_PI426222305 | JHI_St_60k_v1 | DMT400060842 | TAGGCCTGACAGTGGTTACTTGAAGATGAAATGGAAACCCAATGCATGTAGTCTTCCTAG |
Microarray Signals from CUST_23008_PI426222305
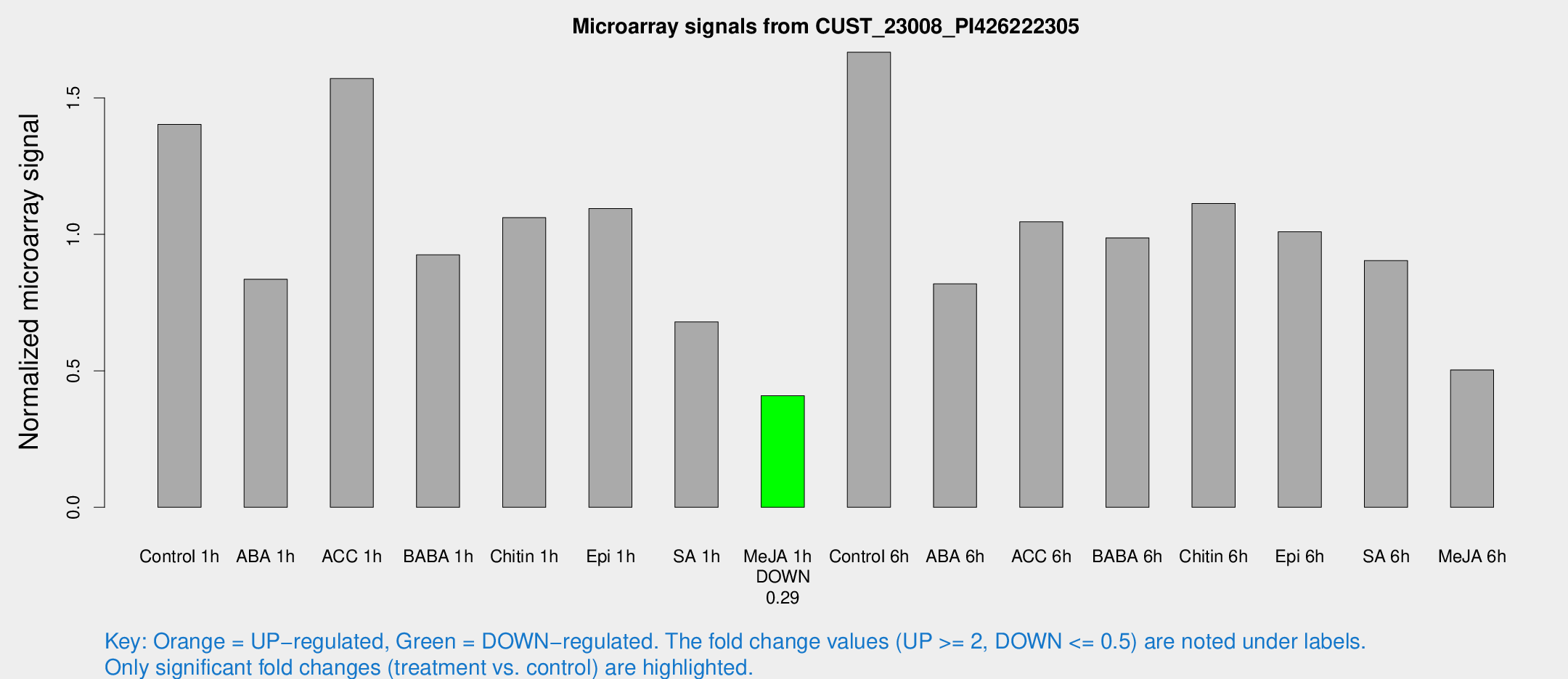
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 27.3338 | 7.93383 | 1.40289 | 0.305268 |
| ABA 1h | 13.6165 | 2.94892 | 0.835623 | 0.189168 |
| ACC 1h | 29.8655 | 5.05873 | 1.57085 | 0.203206 |
| BABA 1h | 17.802 | 5.27638 | 0.924922 | 0.450018 |
| Chitin 1h | 17.6706 | 3.14732 | 1.0614 | 0.197389 |
| Epi 1h | 17.7869 | 3.75443 | 1.09498 | 0.248872 |
| SA 1h | 13.2033 | 3.07716 | 0.679215 | 0.197447 |
| Me-JA 1h | 6.12561 | 3.04374 | 0.409199 | 0.207657 |
| Control 6h | 33.0351 | 8.6991 | 1.66749 | 0.399517 |
| ABA 6h | 15.8404 | 3.33214 | 0.818597 | 0.178201 |
| ACC 6h | 23.1856 | 6.26807 | 1.04609 | 0.189211 |
| BABA 6h | 20.5409 | 3.68336 | 0.987251 | 0.189157 |
| Chitin 6h | 23.4792 | 6.33244 | 1.11361 | 0.364524 |
| Epi 6h | 23.2161 | 7.51312 | 1.00987 | 0.477677 |
| SA 6h | 16.7468 | 3.40454 | 0.904265 | 0.277313 |
| Me-JA 6h | 9.88643 | 3.15493 | 0.50356 | 0.196652 |
Source Transcript PGSC0003DMT400060843 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G06700.1 | +3 | 0.0 | 605 | 280/432 (65%) | Plant protein of unknown function (DUF828) | chr5:2063638-2065810 FORWARD LENGTH=608 |
