Probe CUST_22955_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22955_PI426222305 | JHI_St_60k_v1 | DMT400060842 | TAGGCCTGACAGTGGTTACTTGAAGATGAAATGGAAACCCAATGCATGTAGTCTTCCTAG |
All Microarray Probes Designed to Gene DMG400023668
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_23008_PI426222305 | JHI_St_60k_v1 | DMT400060843 | TAGGCCTGACAGTGGTTACTTGAAGATGAAATGGAAACCCAATGCATGTAGTCTTCCTAG |
| CUST_22955_PI426222305 | JHI_St_60k_v1 | DMT400060842 | TAGGCCTGACAGTGGTTACTTGAAGATGAAATGGAAACCCAATGCATGTAGTCTTCCTAG |
Microarray Signals from CUST_22955_PI426222305
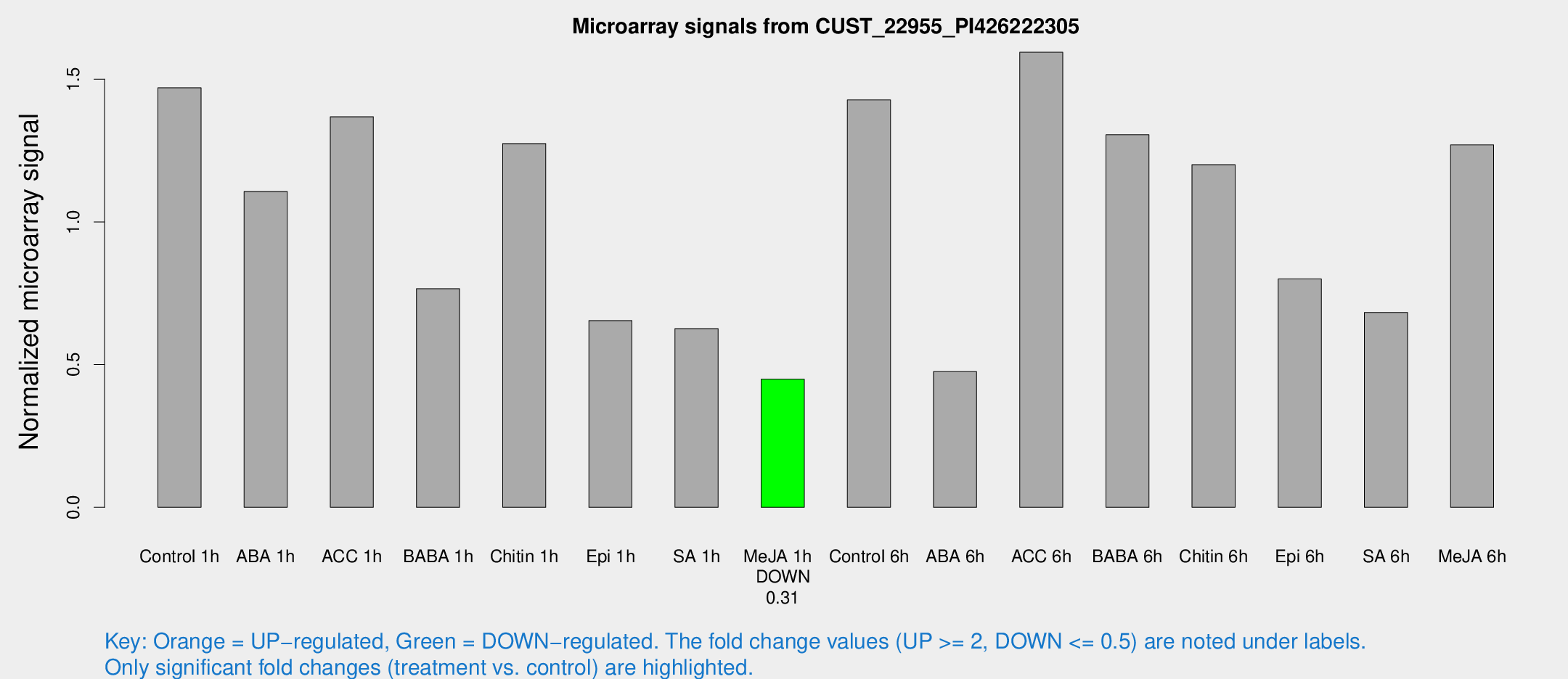
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 23.6986 | 4.57939 | 1.47001 | 0.245809 |
| ABA 1h | 16.8375 | 4.75156 | 1.10667 | 0.536193 |
| ACC 1h | 21.8418 | 3.65854 | 1.36817 | 0.232068 |
| BABA 1h | 11.905 | 3.41623 | 0.766134 | 0.245031 |
| Chitin 1h | 18.4913 | 3.98877 | 1.27453 | 0.341392 |
| Epi 1h | 10.9356 | 5.38844 | 0.65438 | 0.327308 |
| SA 1h | 10.7567 | 3.22816 | 0.625913 | 0.249617 |
| Me-JA 1h | 5.65486 | 3.25753 | 0.448807 | 0.257922 |
| Control 6h | 31.3957 | 14.5136 | 1.42761 | 1.08084 |
| ABA 6h | 8.06972 | 3.43821 | 0.475326 | 0.216452 |
| ACC 6h | 31.4752 | 9.74375 | 1.59453 | 0.385749 |
| BABA 6h | 23.837 | 5.48248 | 1.30526 | 0.271818 |
| Chitin 6h | 21.09 | 5.48238 | 1.20044 | 0.314827 |
| Epi 6h | 14.608 | 3.8914 | 0.800186 | 0.259014 |
| SA 6h | 11.661 | 3.80572 | 0.682682 | 0.361882 |
| Me-JA 6h | 21.0981 | 6.24192 | 1.26977 | 0.283401 |
Source Transcript PGSC0003DMT400060842 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G06700.1 | +3 | 2e-132 | 406 | 190/316 (60%) | Plant protein of unknown function (DUF828) | chr5:2063638-2065810 FORWARD LENGTH=608 |
