Probe CUST_22173_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22173_PI426222305 | JHI_St_60k_v1 | DMT400023517 | GATTCAAGAGCTGTGCTTTCACGTGCTGGAGTTGCAGTACCTTTGTCTATTGATCATAAG |
All Microarray Probes Designed to Gene DMG400009112
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22265_PI426222305 | JHI_St_60k_v1 | DMT400023516 | AAGTCATAAATTGGAATGGACAAAGAGTCTTAGGAGTTCTTGCTACTTCAAGATCCATAG |
| CUST_22173_PI426222305 | JHI_St_60k_v1 | DMT400023517 | GATTCAAGAGCTGTGCTTTCACGTGCTGGAGTTGCAGTACCTTTGTCTATTGATCATAAG |
Microarray Signals from CUST_22173_PI426222305
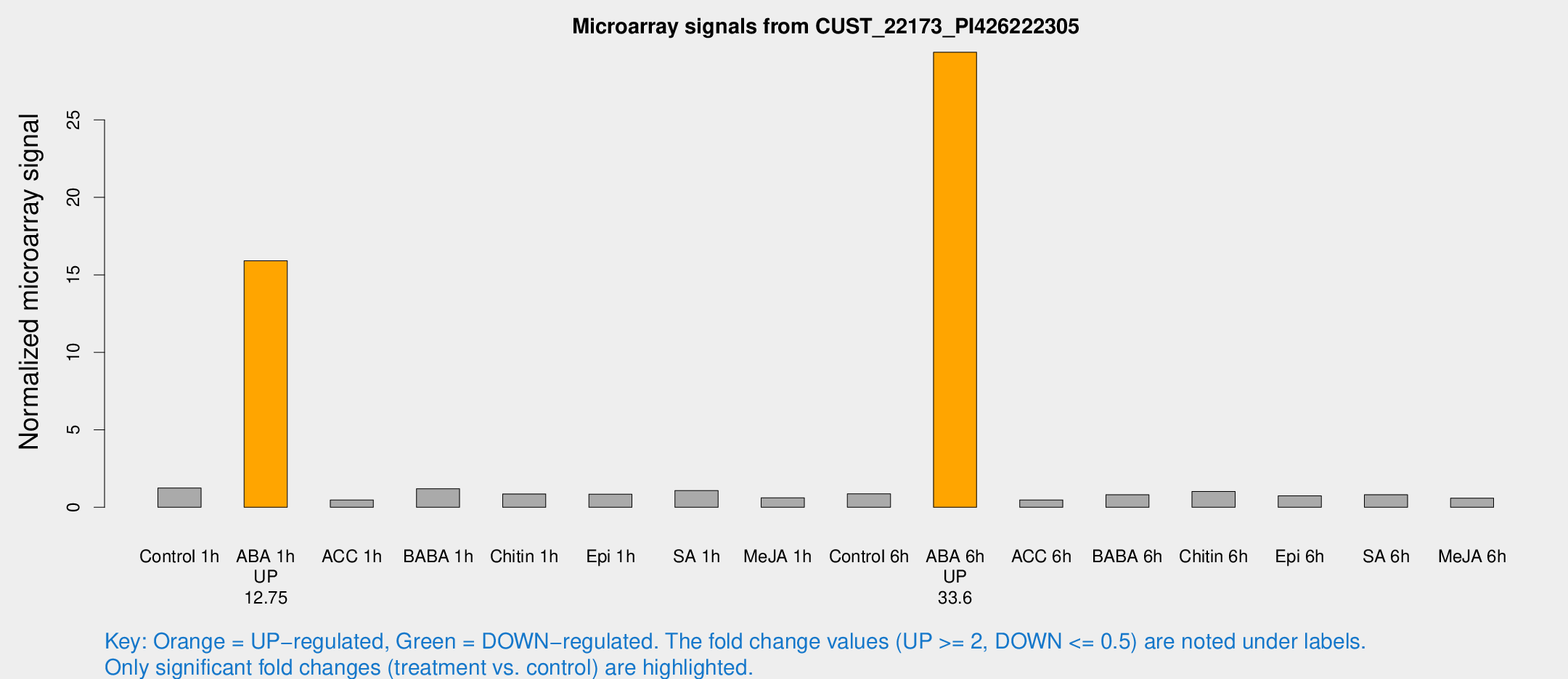
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 115.617 | 12.8371 | 1.24807 | 0.123085 |
| ABA 1h | 1315.54 | 181.517 | 15.9098 | 1.56137 |
| ACC 1h | 82.3456 | 45.1642 | 0.468117 | 0.824195 |
| BABA 1h | 106.324 | 7.24845 | 1.19983 | 0.19939 |
| Chitin 1h | 82.9693 | 34.4554 | 0.862451 | 0.295075 |
| Epi 1h | 71.619 | 15.9883 | 0.849394 | 0.221581 |
| SA 1h | 104.342 | 15.9718 | 1.08077 | 0.251031 |
| Me-JA 1h | 47.3309 | 7.41924 | 0.616919 | 0.133642 |
| Control 6h | 99.6684 | 36.5652 | 0.874092 | 0.432882 |
| ABA 6h | 2882.85 | 266.446 | 29.3665 | 3.33605 |
| ACC 6h | 51.2189 | 7.72458 | 0.477029 | 0.110223 |
| BABA 6h | 84.3535 | 10.8587 | 0.810281 | 0.0728318 |
| Chitin 6h | 108.478 | 31.9562 | 1.01788 | 0.353339 |
| Epi 6h | 78.9082 | 13.8786 | 0.737723 | 0.207361 |
| SA 6h | 82.5758 | 23.9179 | 0.813798 | 0.208815 |
| Me-JA 6h | 75.139 | 43.7051 | 0.59009 | 0.347972 |
Source Transcript PGSC0003DMT400023517 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G11410.1 | +1 | 2e-74 | 244 | 160/360 (44%) | protein phosphatase 2CA | chr3:3584181-3585649 REVERSE LENGTH=399 |
