Probe CUST_22265_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22265_PI426222305 | JHI_St_60k_v1 | DMT400023516 | AAGTCATAAATTGGAATGGACAAAGAGTCTTAGGAGTTCTTGCTACTTCAAGATCCATAG |
All Microarray Probes Designed to Gene DMG400009112
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22265_PI426222305 | JHI_St_60k_v1 | DMT400023516 | AAGTCATAAATTGGAATGGACAAAGAGTCTTAGGAGTTCTTGCTACTTCAAGATCCATAG |
| CUST_22173_PI426222305 | JHI_St_60k_v1 | DMT400023517 | GATTCAAGAGCTGTGCTTTCACGTGCTGGAGTTGCAGTACCTTTGTCTATTGATCATAAG |
Microarray Signals from CUST_22265_PI426222305
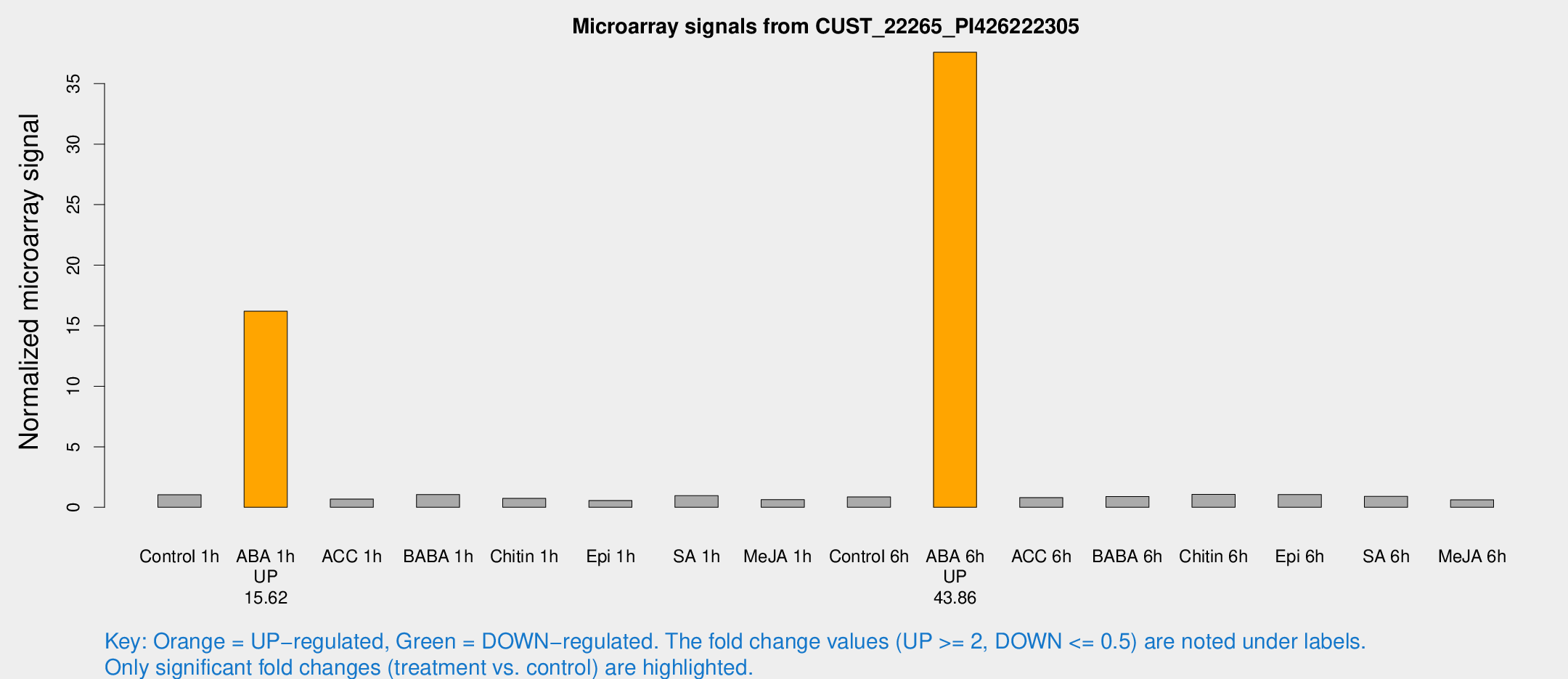
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 48.1693 | 4.41208 | 1.0365 | 0.0950838 |
| ABA 1h | 678.074 | 102.459 | 16.1933 | 1.72709 |
| ACC 1h | 46.2456 | 20.7754 | 0.683241 | 0.559262 |
| BABA 1h | 46.9348 | 4.44799 | 1.05545 | 0.116379 |
| Chitin 1h | 36.6594 | 16.3177 | 0.735618 | 0.27456 |
| Epi 1h | 30.6479 | 12.3538 | 0.561837 | 0.490037 |
| SA 1h | 46.1473 | 5.27523 | 0.960391 | 0.177657 |
| Me-JA 1h | 25.1901 | 4.91743 | 0.638797 | 0.205938 |
| Control 6h | 54.0825 | 21.7464 | 0.856982 | 0.60129 |
| ABA 6h | 1855.02 | 145.64 | 37.5876 | 4.34373 |
| ACC 6h | 42.7362 | 4.88991 | 0.801786 | 0.125221 |
| BABA 6h | 46.9454 | 5.96817 | 0.893331 | 0.0925435 |
| Chitin 6h | 56.5001 | 16.5968 | 1.06439 | 0.308619 |
| Epi 6h | 58.3248 | 14.3767 | 1.05506 | 0.370059 |
| SA 6h | 50.1803 | 21.2184 | 0.898863 | 0.362289 |
| Me-JA 6h | 37.0268 | 15.9755 | 0.618826 | 0.353215 |
Source Transcript PGSC0003DMT400023516 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G11410.1 | +1 | 2e-38 | 141 | 80/146 (55%) | protein phosphatase 2CA | chr3:3584181-3585649 REVERSE LENGTH=399 |
