Probe CUST_22152_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22152_PI426222305 | JHI_St_60k_v1 | DMT400023551 | GCCTCAAATTTTACTGTGATAATTTTGAAGCATTTTTGTGTCATGTTCTTTACCGCTTGG |
All Microarray Probes Designed to Gene DMG400009122
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22213_PI426222305 | JHI_St_60k_v1 | DMT400023552 | CCTTCTGAAATTCAACACTTTGAAGATTTCTATTGCTCACATGAACCAAATGCATCAATG |
| CUST_22152_PI426222305 | JHI_St_60k_v1 | DMT400023551 | GCCTCAAATTTTACTGTGATAATTTTGAAGCATTTTTGTGTCATGTTCTTTACCGCTTGG |
Microarray Signals from CUST_22152_PI426222305
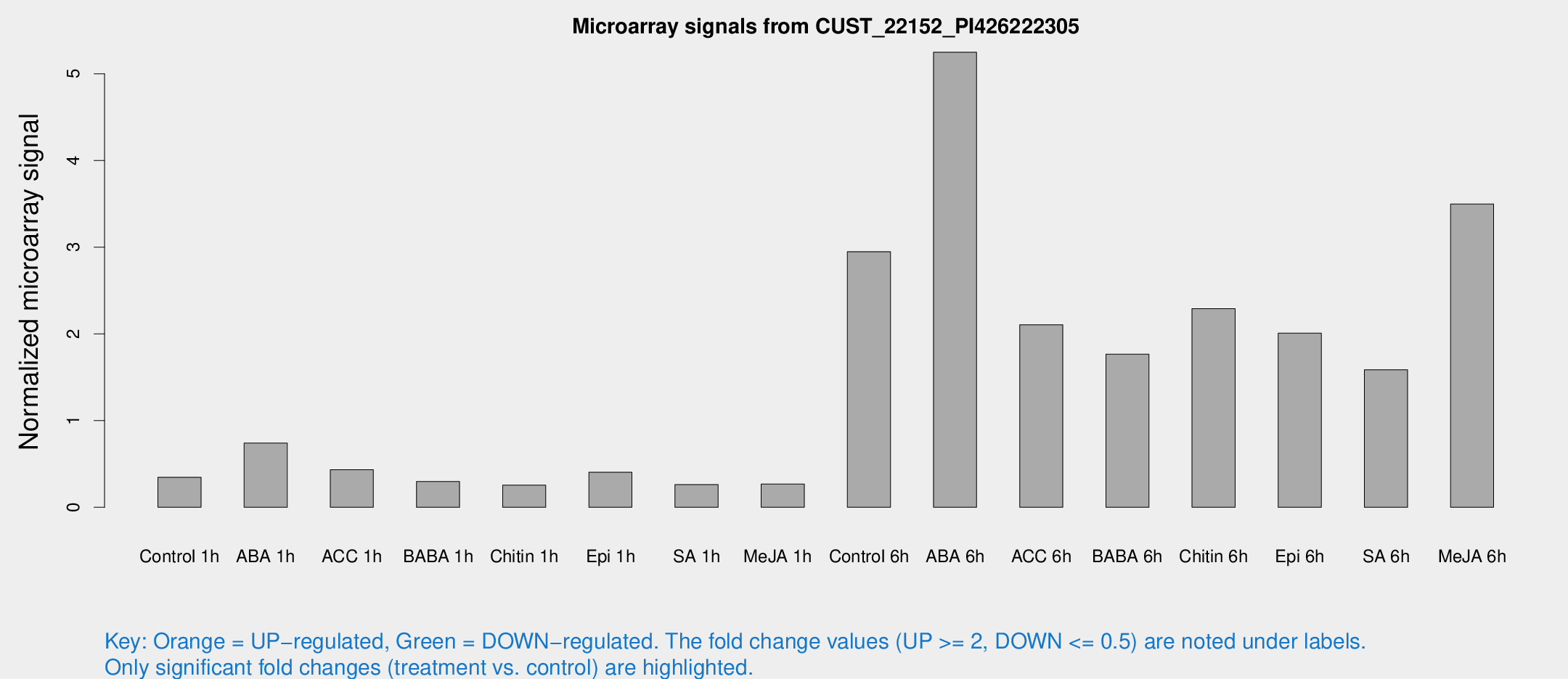
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 11.1177 | 4.59346 | 0.346721 | 0.1762 |
| ABA 1h | 19.6736 | 5.56825 | 0.740992 | 0.182456 |
| ACC 1h | 13.0754 | 4.41169 | 0.433798 | 0.170936 |
| BABA 1h | 8.14227 | 3.73776 | 0.297071 | 0.146986 |
| Chitin 1h | 6.31712 | 3.45364 | 0.255198 | 0.138498 |
| Epi 1h | 10.3605 | 3.38701 | 0.404219 | 0.157123 |
| SA 1h | 7.70131 | 3.4533 | 0.261423 | 0.128542 |
| Me-JA 1h | 6.05323 | 3.53558 | 0.267649 | 0.154972 |
| Control 6h | 85.0608 | 17.0901 | 2.94807 | 0.415293 |
| ABA 6h | 181.494 | 60.5045 | 5.24868 | 2.40289 |
| ACC 6h | 66.5164 | 5.71286 | 2.10548 | 0.27316 |
| BABA 6h | 59.7771 | 19.4014 | 1.76506 | 0.544004 |
| Chitin 6h | 72.2665 | 19.3314 | 2.29117 | 0.719869 |
| Epi 6h | 65.7755 | 13.9964 | 2.0083 | 0.669245 |
| SA 6h | 46.4521 | 12.8847 | 1.58628 | 0.68443 |
| Me-JA 6h | 117.409 | 46.5778 | 3.49762 | 1.57519 |
Source Transcript PGSC0003DMT400023551 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G53420.1 | +2 | 6e-57 | 196 | 124/239 (52%) | CCT motif family protein | chr5:21673683-21675469 FORWARD LENGTH=264 |
