Probe CUST_22213_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22213_PI426222305 | JHI_St_60k_v1 | DMT400023552 | CCTTCTGAAATTCAACACTTTGAAGATTTCTATTGCTCACATGAACCAAATGCATCAATG |
All Microarray Probes Designed to Gene DMG400009122
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22213_PI426222305 | JHI_St_60k_v1 | DMT400023552 | CCTTCTGAAATTCAACACTTTGAAGATTTCTATTGCTCACATGAACCAAATGCATCAATG |
| CUST_22152_PI426222305 | JHI_St_60k_v1 | DMT400023551 | GCCTCAAATTTTACTGTGATAATTTTGAAGCATTTTTGTGTCATGTTCTTTACCGCTTGG |
Microarray Signals from CUST_22213_PI426222305
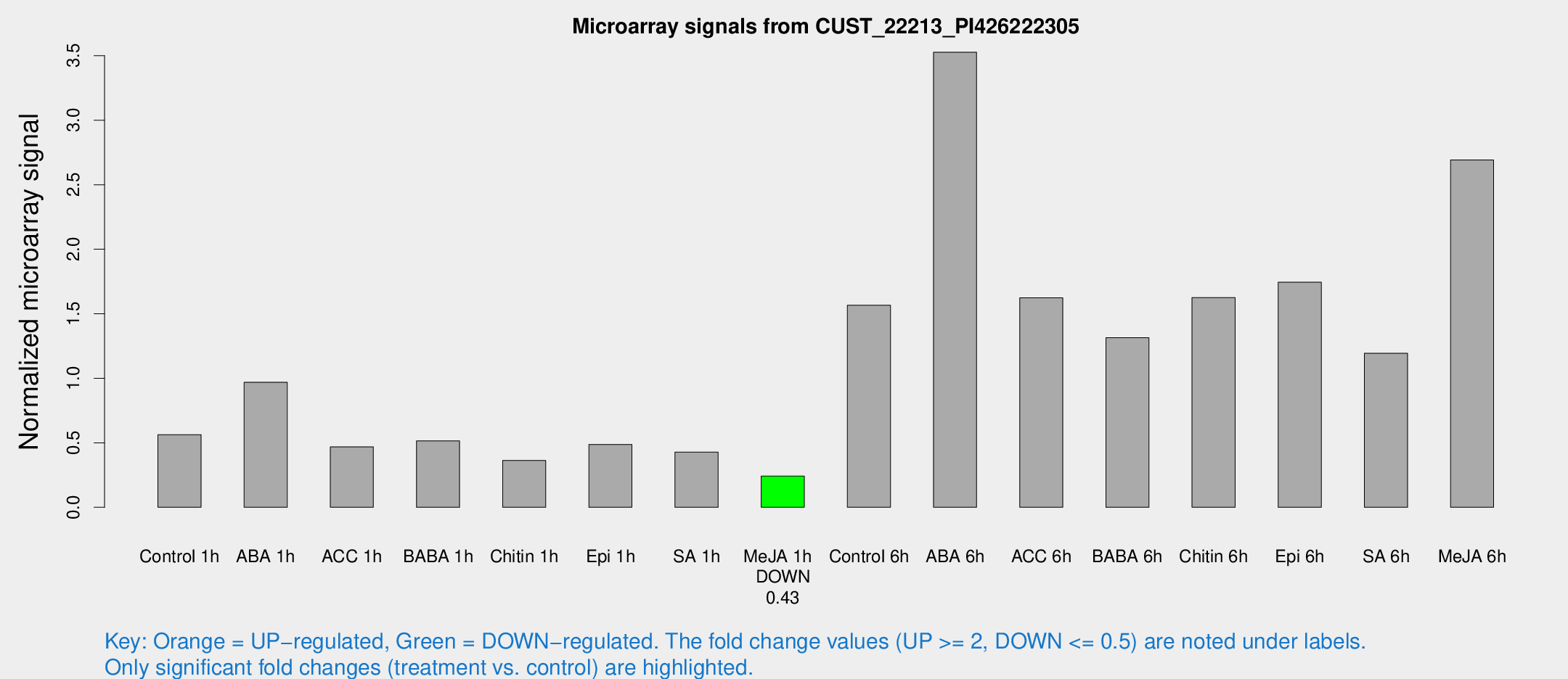
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 113.018 | 15.5365 | 0.563268 | 0.0393471 |
| ABA 1h | 169.36 | 10.1969 | 0.969142 | 0.0582565 |
| ACC 1h | 106.104 | 30.3694 | 0.468036 | 0.141828 |
| BABA 1h | 102.253 | 21.0181 | 0.515462 | 0.0659756 |
| Chitin 1h | 66.7534 | 13.2803 | 0.363239 | 0.0475232 |
| Epi 1h | 86.2875 | 15.5503 | 0.487097 | 0.0947232 |
| SA 1h | 90.1579 | 17.4624 | 0.427282 | 0.0752746 |
| Me-JA 1h | 39.4442 | 4.16635 | 0.242198 | 0.0235011 |
| Control 6h | 318.422 | 52.8232 | 1.56612 | 0.165288 |
| ABA 6h | 813.104 | 216.506 | 3.52684 | 1.06645 |
| ACC 6h | 369.298 | 29.1435 | 1.62373 | 0.0992329 |
| BABA 6h | 296.656 | 46.112 | 1.31489 | 0.188761 |
| Chitin 6h | 349.982 | 57.6583 | 1.6257 | 0.301841 |
| Epi 6h | 391.57 | 37.8595 | 1.74445 | 0.300276 |
| SA 6h | 244.409 | 56.5754 | 1.19307 | 0.204457 |
| Me-JA 6h | 584.006 | 179.145 | 2.6916 | 0.665299 |
Source Transcript PGSC0003DMT400023552 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G53420.1 | +3 | 4e-67 | 219 | 139/279 (50%) | CCT motif family protein | chr5:21673683-21675469 FORWARD LENGTH=264 |
