Probe CUST_2184_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_2184_PI426222305 | JHI_St_60k_v1 | DMT400072556 | CATCAGTAACTGATTCCGAGGCATCGCATGATGGTTGGAAAATGAAGGACAATGGTAACA |
All Microarray Probes Designed to Gene DMG400028231
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_2184_PI426222305 | JHI_St_60k_v1 | DMT400072556 | CATCAGTAACTGATTCCGAGGCATCGCATGATGGTTGGAAAATGAAGGACAATGGTAACA |
| CUST_2542_PI426222305 | JHI_St_60k_v1 | DMT400072557 | GGCTTTCATGAATTTTAGAGGACTGTTTACTTTTTCTTGGAGATGCAATATGAAGGACAA |
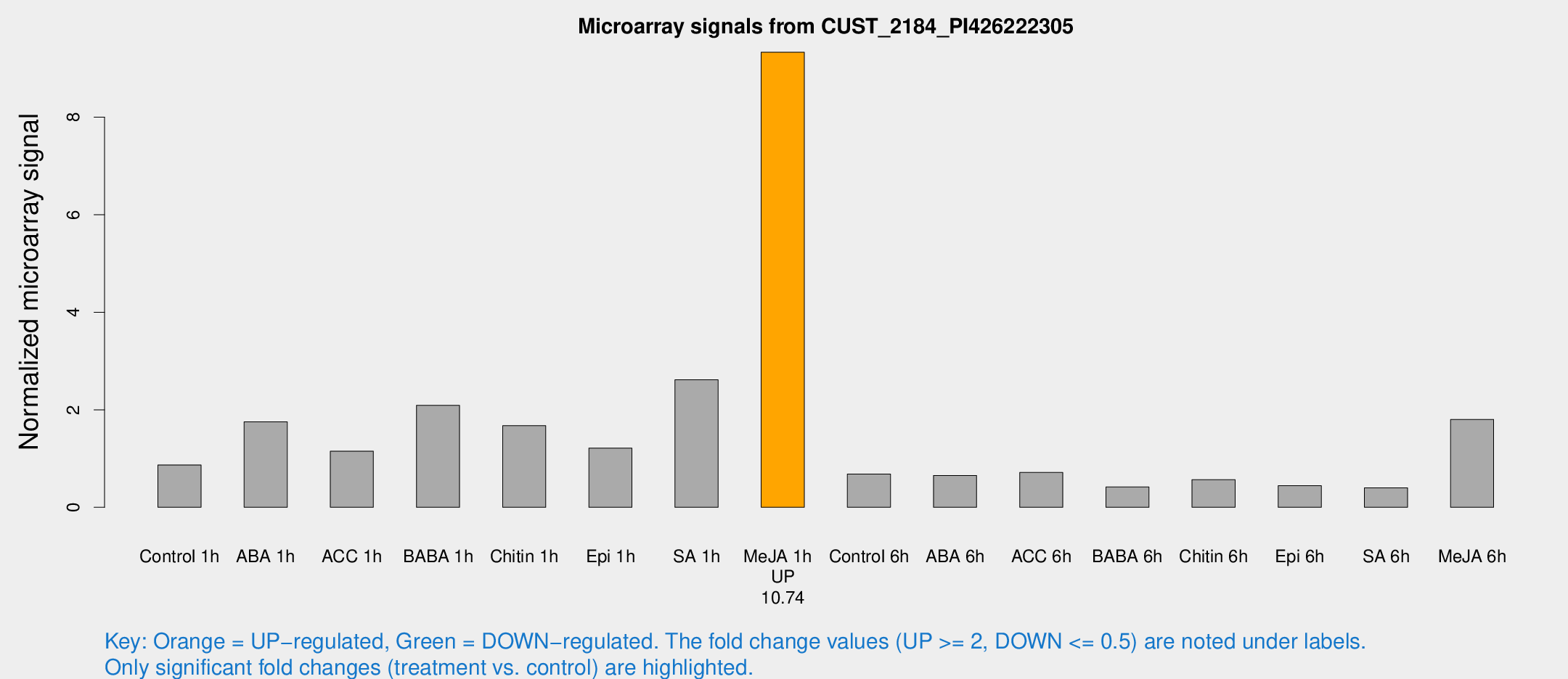
Microarray Signals from CUST_2184_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 25.8265 | 10.4429 | 0.869053 | 0.585464 |
| ABA 1h | 37.1086 | 6.64377 | 1.75171 | 0.192249 |
| ACC 1h | 45.1551 | 29.0187 | 1.15093 | 1.13727 |
| BABA 1h | 48.4691 | 9.45052 | 2.08981 | 0.644556 |
| Chitin 1h | 39.9833 | 14.8982 | 1.67324 | 0.74916 |
| Epi 1h | 26.3317 | 6.5439 | 1.21455 | 0.356905 |
| SA 1h | 77.4195 | 33.831 | 2.61515 | 1.65303 |
| Me-JA 1h | 184.683 | 37.3541 | 9.33284 | 1.14367 |
| Control 6h | 18.3065 | 5.69688 | 0.681148 | 0.235252 |
| ABA 6h | 19.1758 | 6.40702 | 0.653771 | 0.309937 |
| ACC 6h | 21.8003 | 6.70008 | 0.714035 | 0.284698 |
| BABA 6h | 10.8742 | 4.35219 | 0.416847 | 0.165983 |
| Chitin 6h | 14.6856 | 4.31144 | 0.56471 | 0.183792 |
| Epi 6h | 12.6559 | 4.65692 | 0.443882 | 0.191774 |
| SA 6h | 9.48437 | 4.08816 | 0.399628 | 0.184309 |
| Me-JA 6h | 41.8411 | 4.48083 | 1.80243 | 0.192667 |
Source Transcript PGSC0003DMT400072556 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | Solyc10g081770.1 | +1 | 0.0 | 843 | 435/472 (92%) | evidence_code:10F0H1E0IEG genomic_reference:SL2.50ch10 gene_region:62095810-62099028 transcript_region:SL2.50ch10:62095810..62099028- functional_description:cDNA clone J023119C16 full insert sequence (AHRD V1 *-*- B7F733_ORYSJ) |
| TAIR PP10 | AT5G06440.4 | +1 | 6e-124 | 374 | 207/439 (47%) | FUNCTIONS IN: molecular_function unknown; INVOLVED IN: biological_process unknown; LOCATED IN: mitochondrion; EXPRESSED IN: 22 plant structures; EXPRESSED DURING: 12 growth stages; BEST Arabidopsis thaliana protein match is: Polyketide cyclase/dehydrase and lipid transport superfamily protein (TAIR:AT3G11720.3). | chr5:1964641-1966807 REVERSE LENGTH=486 |
