Probe CUST_21287_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_21287_PI426222305 | JHI_St_60k_v1 | DMT400020222 | CCTAGATACGTAAGGATATCATGTAGCTATGGTGAAGGAGATGGTAACTAAATATTACAT |
All Microarray Probes Designed to Gene DMG400007815
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_21147_PI426222305 | JHI_St_60k_v1 | DMT400020220 | ATATAATTTTATAGCAGGCCTTATCCACAATCCTCAACTACATGAGGGTCGGGTGGCACG |
| CUST_21287_PI426222305 | JHI_St_60k_v1 | DMT400020222 | CCTAGATACGTAAGGATATCATGTAGCTATGGTGAAGGAGATGGTAACTAAATATTACAT |
Microarray Signals from CUST_21287_PI426222305
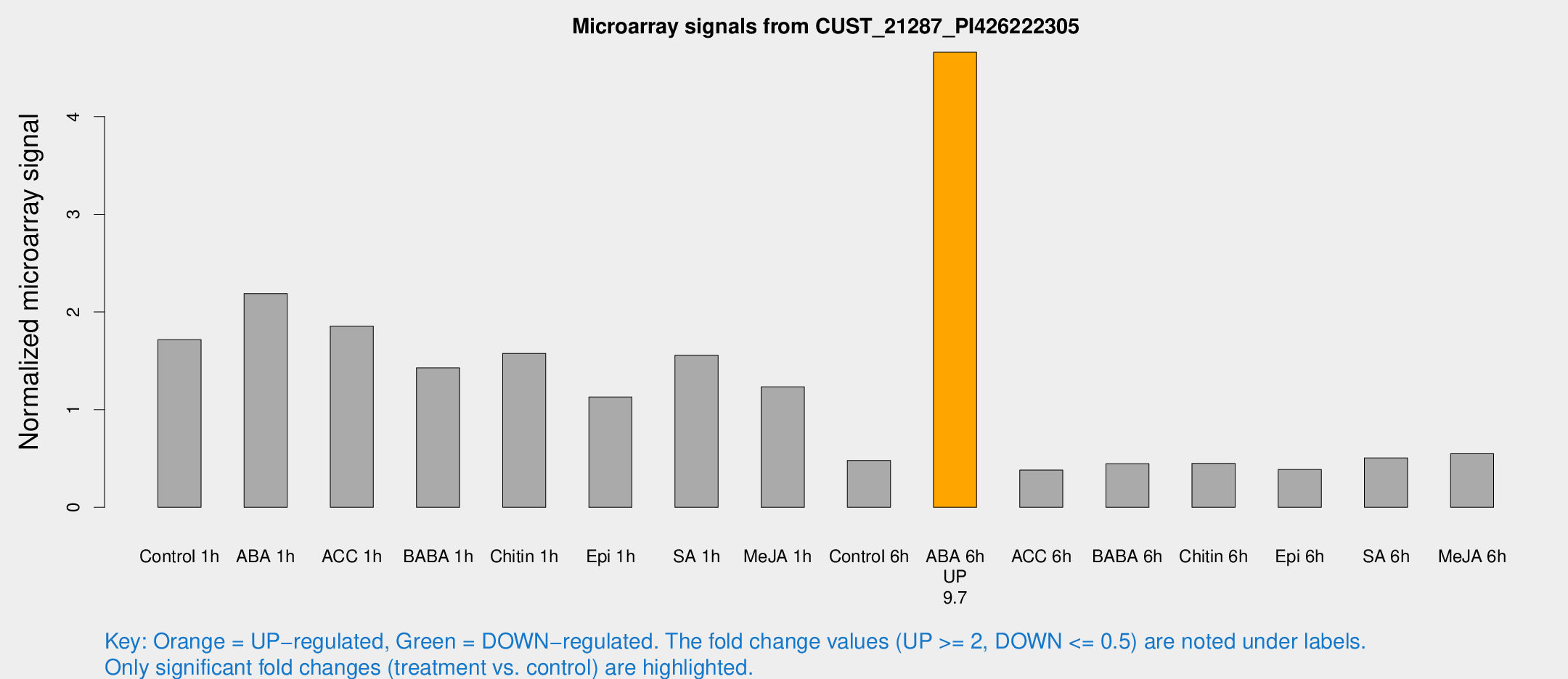
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 3719.44 | 953.386 | 1.71619 | 0.344448 |
| ABA 1h | 4290.82 | 1270.19 | 2.18911 | 0.844325 |
| ACC 1h | 3899.59 | 225.92 | 1.85576 | 0.253994 |
| BABA 1h | 2828.85 | 235.821 | 1.42838 | 0.255734 |
| Chitin 1h | 2954.5 | 424.967 | 1.5759 | 0.186692 |
| Epi 1h | 2021.34 | 229.576 | 1.12894 | 0.120644 |
| SA 1h | 3467.65 | 770.73 | 1.55715 | 0.520554 |
| Me-JA 1h | 2354.84 | 889.011 | 1.23348 | 0.55932 |
| Control 6h | 1123.35 | 374.426 | 0.480539 | 0.14862 |
| ABA 6h | 10390.1 | 1616.96 | 4.66006 | 0.860593 |
| ACC 6h | 891.934 | 51.7194 | 0.380681 | 0.0514606 |
| BABA 6h | 1107.3 | 335.079 | 0.446305 | 0.152787 |
| Chitin 6h | 1039.75 | 265.677 | 0.449626 | 0.124555 |
| Epi 6h | 891.45 | 51.7284 | 0.386737 | 0.0465901 |
| SA 6h | 1059.87 | 212.345 | 0.506316 | 0.0799389 |
| Me-JA 6h | 1160.39 | 247.079 | 0.548157 | 0.0790041 |
Source Transcript PGSC0003DMT400020222 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G30450.1 | +1 | 7e-06 | 44 | 20/35 (57%) | glycine-rich protein | chr4:14886283-14886606 REVERSE LENGTH=107 |
