Probe CUST_21147_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_21147_PI426222305 | JHI_St_60k_v1 | DMT400020220 | ATATAATTTTATAGCAGGCCTTATCCACAATCCTCAACTACATGAGGGTCGGGTGGCACG |
All Microarray Probes Designed to Gene DMG400007815
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_21147_PI426222305 | JHI_St_60k_v1 | DMT400020220 | ATATAATTTTATAGCAGGCCTTATCCACAATCCTCAACTACATGAGGGTCGGGTGGCACG |
| CUST_21287_PI426222305 | JHI_St_60k_v1 | DMT400020222 | CCTAGATACGTAAGGATATCATGTAGCTATGGTGAAGGAGATGGTAACTAAATATTACAT |
Microarray Signals from CUST_21147_PI426222305
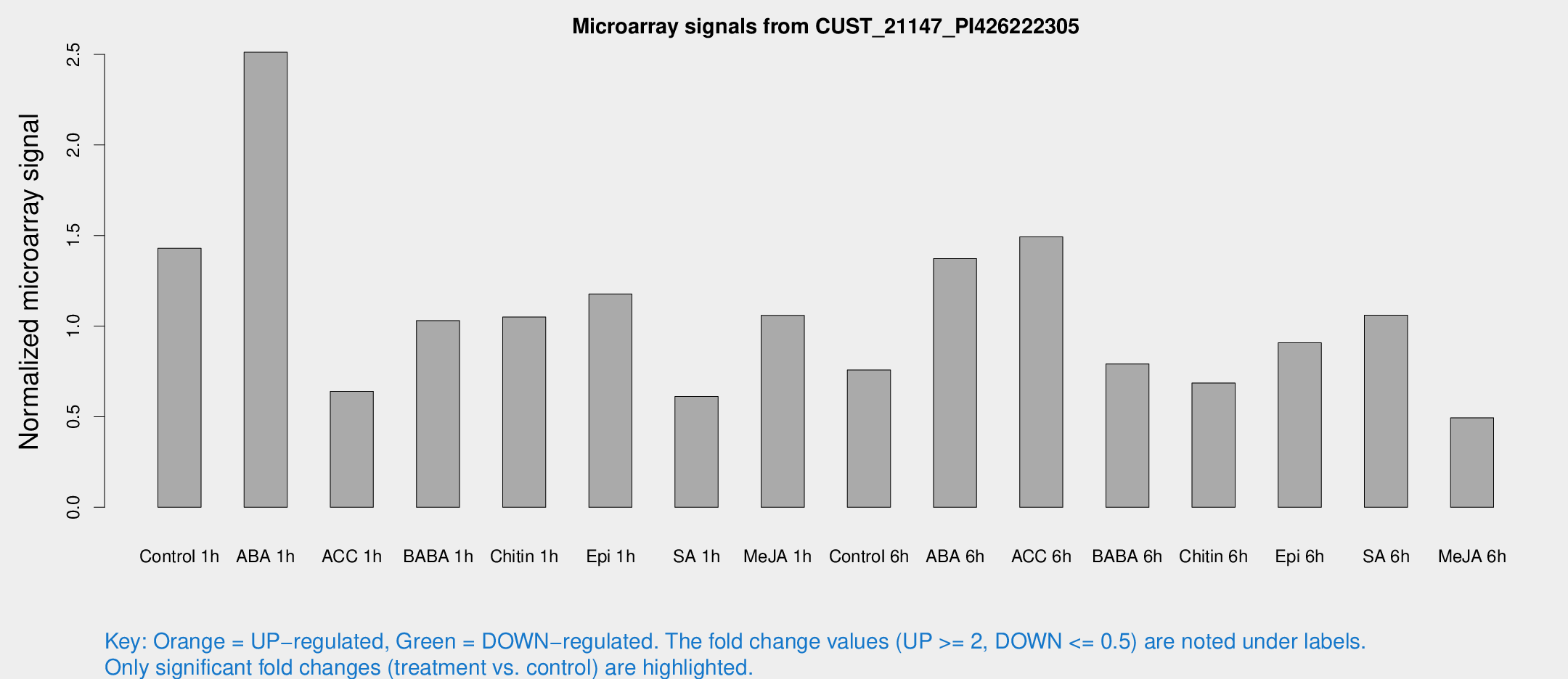
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 22.0647 | 3.94153 | 1.42942 | 0.256672 |
| ABA 1h | 35.1577 | 5.67513 | 2.51185 | 0.325717 |
| ACC 1h | 10.8791 | 4.32897 | 0.640267 | 0.291126 |
| BABA 1h | 17.6523 | 5.46064 | 1.03079 | 0.33643 |
| Chitin 1h | 16.491 | 6.12627 | 1.04995 | 0.407676 |
| Epi 1h | 20.2216 | 8.4251 | 1.17715 | 0.795764 |
| SA 1h | 9.88508 | 3.66251 | 0.612014 | 0.234905 |
| Me-JA 1h | 13.6754 | 3.85105 | 1.0597 | 0.309326 |
| Control 6h | 13.7725 | 5.75638 | 0.757792 | 0.306302 |
| ABA 6h | 22.5293 | 4.19792 | 1.37267 | 0.256 |
| ACC 6h | 26.6775 | 4.85047 | 1.49269 | 0.270048 |
| BABA 6h | 20.6318 | 13.2372 | 0.79152 | 0.656062 |
| Chitin 6h | 12.111 | 4.32227 | 0.686173 | 0.283094 |
| Epi 6h | 16.2862 | 4.89945 | 0.908078 | 0.290606 |
| SA 6h | 17.1458 | 4.07893 | 1.06049 | 0.296685 |
| Me-JA 6h | 7.76735 | 3.83496 | 0.493852 | 0.254728 |
Source Transcript PGSC0003DMT400020220 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G31540.1 | +1 | 3e-65 | 213 | 101/202 (50%) | GDSL-like Lipase/Acylhydrolase superfamily protein | chr2:13430733-13432045 REVERSE LENGTH=360 |
