Probe CUST_20813_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_20813_PI426222305 | JHI_St_60k_v1 | DMT400011726 | TGGGTTGAATCGAATCGAGAGGTCAAGTTTGAATCATTCTTCTGTTGTCCGTTATTTTTT |
All Microarray Probes Designed to Gene DMG400004601
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_20712_PI426222305 | JHI_St_60k_v1 | DMT400011723 | TCTTTTGCTATTAAGATTTTGGAGAAGAATCGGATCCAGGACCTTCGAATCACTGATCAG |
| CUST_20763_PI426222305 | JHI_St_60k_v1 | DMT400011720 | TGTGTTTAACTTGTTTTTGTGCACATAGAGAGATATCTACCCTAAAACACTAGCTCCCAT |
| CUST_20834_PI426222305 | JHI_St_60k_v1 | DMT400011725 | TCTTTTGCTATTAAGATTTTGGAGAAGAATCGGATCCAGGACCTTCGAATCACTGATCAG |
| CUST_20913_PI426222305 | JHI_St_60k_v1 | DMT400011724 | TCTTTTGCTATTAAGATTTTGGAGAAGAATCGGATCCAGGACCTTCGAATCACTGATCAG |
| CUST_20957_PI426222305 | JHI_St_60k_v1 | DMT400011719 | TCTTTTGCTATTAAGATTTTGGAGAAGAATCGGATCCAGGACCTTCGAATCACTGATCAG |
| CUST_20663_PI426222305 | JHI_St_60k_v1 | DMT400011718 | ACATCTTAGGGTATGGTACTTAACTTGTGACTAGTAATAGTCGAGGCGAATCTAACGTTT |
| CUST_20813_PI426222305 | JHI_St_60k_v1 | DMT400011726 | TGGGTTGAATCGAATCGAGAGGTCAAGTTTGAATCATTCTTCTGTTGTCCGTTATTTTTT |
| CUST_20681_PI426222305 | JHI_St_60k_v1 | DMT400011721 | ATAGTCCTAAGAAATTCCTGAGGGCCAGTAACACACGGTTCGATTCTTGGTGGATAATGG |
Microarray Signals from CUST_20813_PI426222305
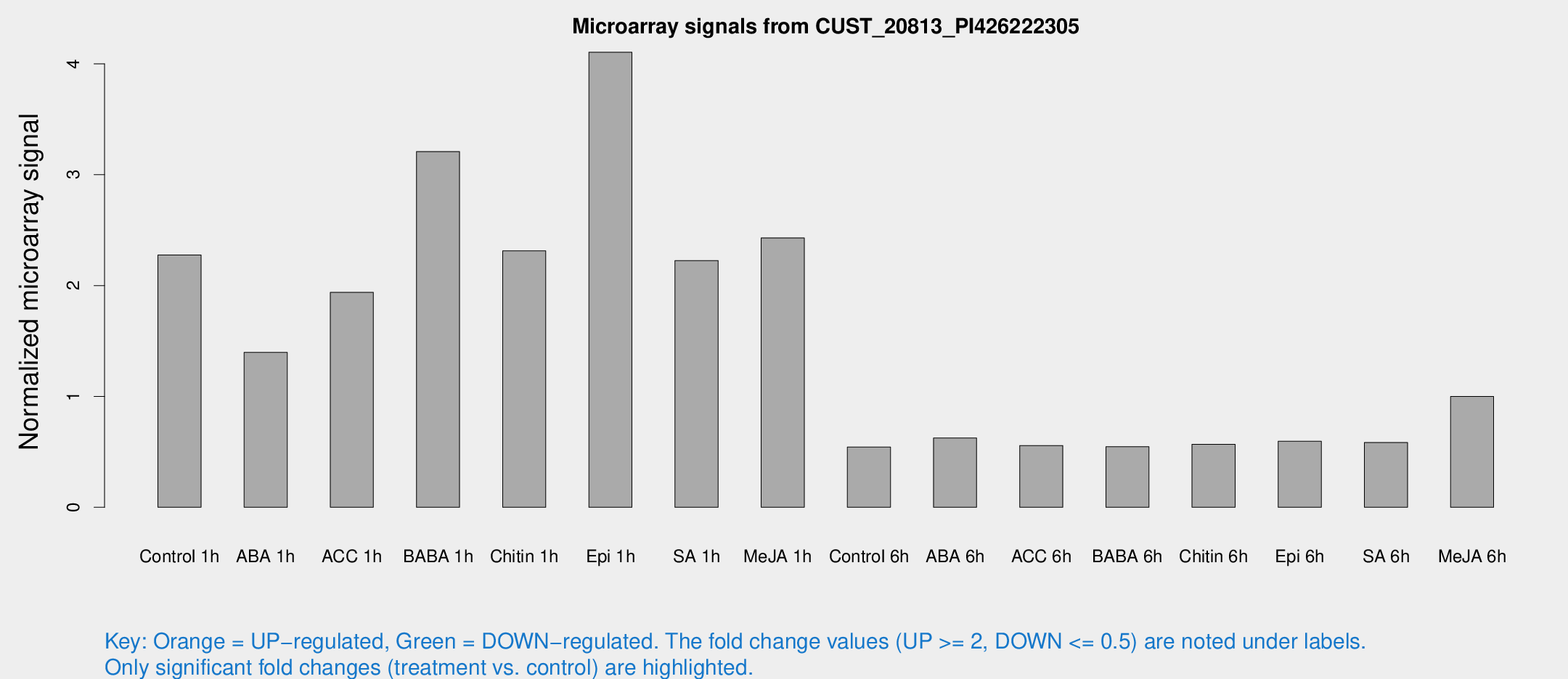
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 24.9132 | 10.2494 | 2.27588 | 0.812806 |
| ABA 1h | 13.6106 | 4.73301 | 1.39667 | 0.824555 |
| ACC 1h | 22.4523 | 8.29164 | 1.93837 | 1.1735 |
| BABA 1h | 32.4526 | 9.19483 | 3.20879 | 0.860184 |
| Chitin 1h | 21.326 | 6.30531 | 2.31271 | 0.531319 |
| Epi 1h | 34.4501 | 6.53948 | 4.1046 | 0.642513 |
| SA 1h | 23.8904 | 7.84141 | 2.22592 | 0.712747 |
| Me-JA 1h | 20.2256 | 5.49854 | 2.43 | 0.71529 |
| Control 6h | 5.10534 | 2.96143 | 0.543504 | 0.315007 |
| ABA 6h | 6.42932 | 3.09484 | 0.624796 | 0.31788 |
| ACC 6h | 6.11753 | 3.62134 | 0.557142 | 0.322793 |
| BABA 6h | 5.74694 | 3.33552 | 0.546371 | 0.316432 |
| Chitin 6h | 5.6757 | 3.28842 | 0.568462 | 0.329267 |
| Epi 6h | 6.32072 | 3.44195 | 0.595766 | 0.322677 |
| SA 6h | 5.4192 | 3.14099 | 0.583909 | 0.33822 |
| Me-JA 6h | 10.2598 | 3.05614 | 0.999319 | 0.366235 |
Source Transcript PGSC0003DMT400011726 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G17510.1 | +2 | 1e-17 | 82 | 41/65 (63%) | CBL-interacting protein kinase 1 | chr3:5989309-5992627 REVERSE LENGTH=444 |
