Probe CUST_20681_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_20681_PI426222305 | JHI_St_60k_v1 | DMT400011721 | ATAGTCCTAAGAAATTCCTGAGGGCCAGTAACACACGGTTCGATTCTTGGTGGATAATGG |
All Microarray Probes Designed to Gene DMG400004601
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_20712_PI426222305 | JHI_St_60k_v1 | DMT400011723 | TCTTTTGCTATTAAGATTTTGGAGAAGAATCGGATCCAGGACCTTCGAATCACTGATCAG |
| CUST_20763_PI426222305 | JHI_St_60k_v1 | DMT400011720 | TGTGTTTAACTTGTTTTTGTGCACATAGAGAGATATCTACCCTAAAACACTAGCTCCCAT |
| CUST_20834_PI426222305 | JHI_St_60k_v1 | DMT400011725 | TCTTTTGCTATTAAGATTTTGGAGAAGAATCGGATCCAGGACCTTCGAATCACTGATCAG |
| CUST_20913_PI426222305 | JHI_St_60k_v1 | DMT400011724 | TCTTTTGCTATTAAGATTTTGGAGAAGAATCGGATCCAGGACCTTCGAATCACTGATCAG |
| CUST_20957_PI426222305 | JHI_St_60k_v1 | DMT400011719 | TCTTTTGCTATTAAGATTTTGGAGAAGAATCGGATCCAGGACCTTCGAATCACTGATCAG |
| CUST_20663_PI426222305 | JHI_St_60k_v1 | DMT400011718 | ACATCTTAGGGTATGGTACTTAACTTGTGACTAGTAATAGTCGAGGCGAATCTAACGTTT |
| CUST_20813_PI426222305 | JHI_St_60k_v1 | DMT400011726 | TGGGTTGAATCGAATCGAGAGGTCAAGTTTGAATCATTCTTCTGTTGTCCGTTATTTTTT |
| CUST_20681_PI426222305 | JHI_St_60k_v1 | DMT400011721 | ATAGTCCTAAGAAATTCCTGAGGGCCAGTAACACACGGTTCGATTCTTGGTGGATAATGG |
Microarray Signals from CUST_20681_PI426222305
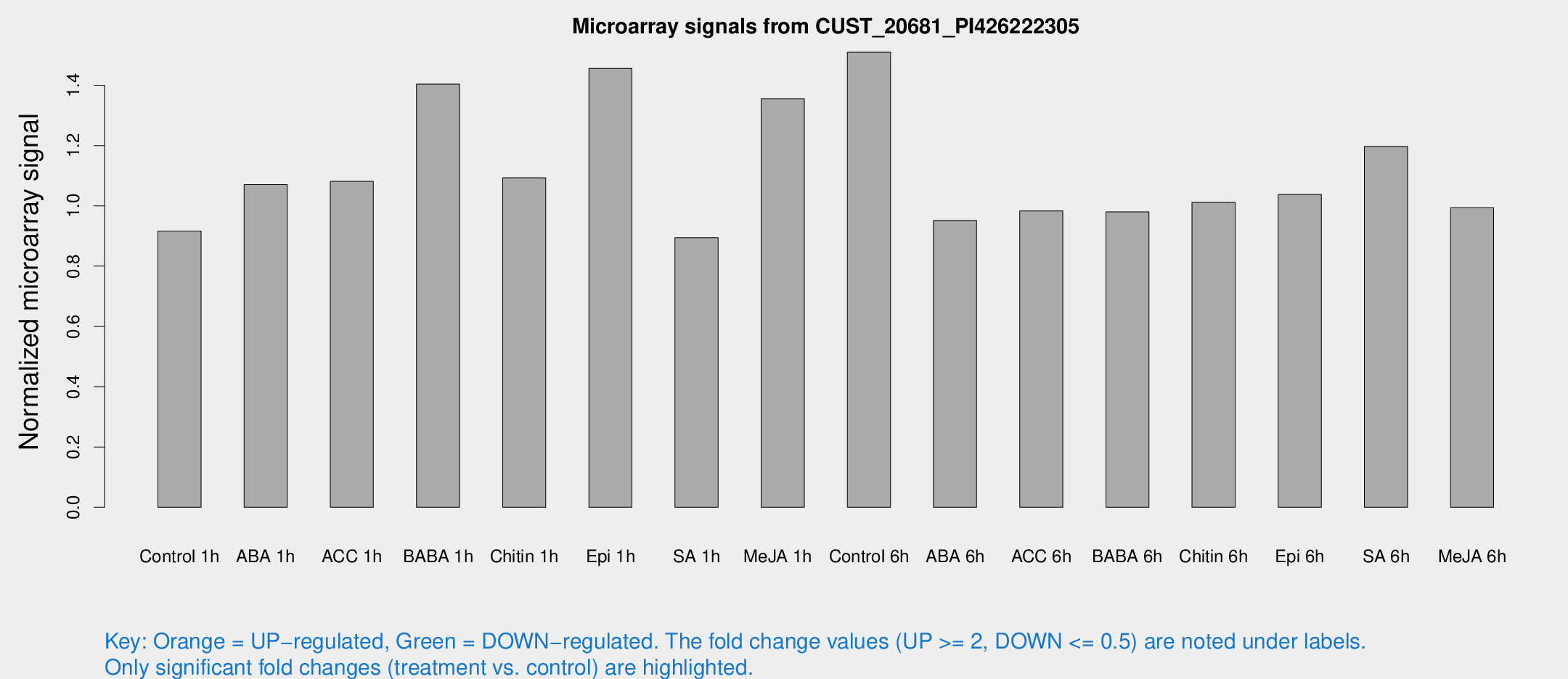
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 5.39183 | 3.12602 | 0.916147 | 0.530497 |
| ABA 1h | 5.60204 | 3.07052 | 1.07064 | 0.591264 |
| ACC 1h | 6.61628 | 3.44883 | 1.08141 | 0.576085 |
| BABA 1h | 8.79351 | 3.35601 | 1.40395 | 0.662842 |
| Chitin 1h | 5.79006 | 3.21736 | 1.09322 | 0.603078 |
| Epi 1h | 8.03364 | 3.14379 | 1.45578 | 0.686939 |
| SA 1h | 5.40578 | 3.13474 | 0.894184 | 0.517895 |
| Me-JA 1h | 6.60601 | 3.27481 | 1.35526 | 0.693172 |
| Control 6h | 9.30458 | 3.42198 | 1.50955 | 0.62166 |
| ABA 6h | 5.94894 | 3.44655 | 0.951472 | 0.550959 |
| ACC 6h | 6.7707 | 4.01156 | 0.983165 | 0.569612 |
| BABA 6h | 6.4699 | 3.76597 | 0.980031 | 0.56862 |
| Chitin 6h | 6.33029 | 3.66723 | 1.01121 | 0.585515 |
| Epi 6h | 6.95243 | 3.82424 | 1.03783 | 0.56714 |
| SA 6h | 7.02423 | 3.52385 | 1.19682 | 0.611181 |
| Me-JA 6h | 5.82209 | 3.37547 | 0.99353 | 0.575355 |
Source Transcript PGSC0003DMT400011721 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G48260.1 | +1 | 1e-16 | 83 | 35/55 (64%) | CBL-interacting protein kinase 17 | chr1:17814226-17817226 REVERSE LENGTH=432 |
