Probe CUST_20357_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_20357_PI426222305 | JHI_St_60k_v1 | DMT400049820 | TGGTGTTAAACTCTTTGCTCTCAAGGTTAATCTTCTCTTCTTTAGATTCATGGGCTTCAT |
All Microarray Probes Designed to Gene DMG401019352
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_20357_PI426222305 | JHI_St_60k_v1 | DMT400049820 | TGGTGTTAAACTCTTTGCTCTCAAGGTTAATCTTCTCTTCTTTAGATTCATGGGCTTCAT |
| CUST_20351_PI426222305 | JHI_St_60k_v1 | DMT400049817 | AACAAACAAATTAATCAACTGTGACGCTTCTTTGCAGCATGTCGTGAAAGCACCAGCTAA |
Microarray Signals from CUST_20357_PI426222305
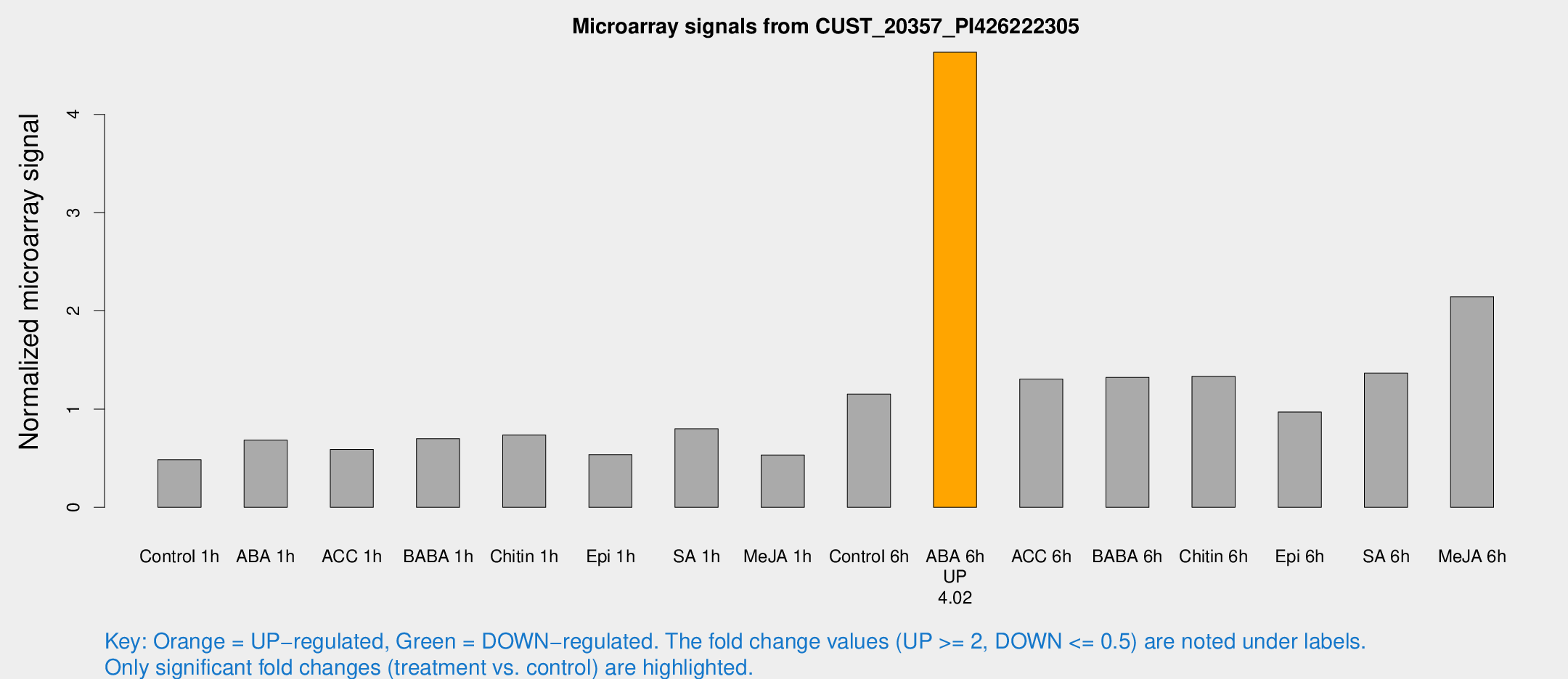
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 67.5788 | 11.6339 | 0.483939 | 0.0645726 |
| ABA 1h | 88.8888 | 23.4236 | 0.68306 | 0.191271 |
| ACC 1h | 87.9425 | 25.2138 | 0.589013 | 0.14631 |
| BABA 1h | 101.311 | 28.7561 | 0.698869 | 0.181169 |
| Chitin 1h | 99.3611 | 34.0305 | 0.735975 | 0.188305 |
| Epi 1h | 66.3883 | 14.9178 | 0.536027 | 0.128127 |
| SA 1h | 127.909 | 50.8062 | 0.799898 | 0.268016 |
| Me-JA 1h | 62.5755 | 16.3528 | 0.532818 | 0.0896349 |
| Control 6h | 160.088 | 26.4129 | 1.1528 | 0.107275 |
| ABA 6h | 689.176 | 134.874 | 4.63274 | 0.623383 |
| ACC 6h | 202.792 | 12.3074 | 1.30528 | 0.16947 |
| BABA 6h | 200.677 | 12.8715 | 1.32299 | 0.0795891 |
| Chitin 6h | 200.379 | 38.7224 | 1.33362 | 0.265305 |
| Epi 6h | 156.757 | 37.2392 | 0.970245 | 0.170237 |
| SA 6h | 187.63 | 29.7782 | 1.36692 | 0.0890139 |
| Me-JA 6h | 294.399 | 44.2761 | 2.14484 | 0.176064 |
Source Transcript PGSC0003DMT400049820 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G03090.2 | +1 | 9e-39 | 141 | 66/126 (52%) | methylcrotonyl-CoA carboxylase alpha chain, mitochondrial / 3-methylcrotonyl-CoA carboxylase 1 (MCCA) | chr1:739715-743819 FORWARD LENGTH=734 |
