Probe CUST_20351_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_20351_PI426222305 | JHI_St_60k_v1 | DMT400049817 | AACAAACAAATTAATCAACTGTGACGCTTCTTTGCAGCATGTCGTGAAAGCACCAGCTAA |
All Microarray Probes Designed to Gene DMG401019352
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_20357_PI426222305 | JHI_St_60k_v1 | DMT400049820 | TGGTGTTAAACTCTTTGCTCTCAAGGTTAATCTTCTCTTCTTTAGATTCATGGGCTTCAT |
| CUST_20351_PI426222305 | JHI_St_60k_v1 | DMT400049817 | AACAAACAAATTAATCAACTGTGACGCTTCTTTGCAGCATGTCGTGAAAGCACCAGCTAA |
Microarray Signals from CUST_20351_PI426222305
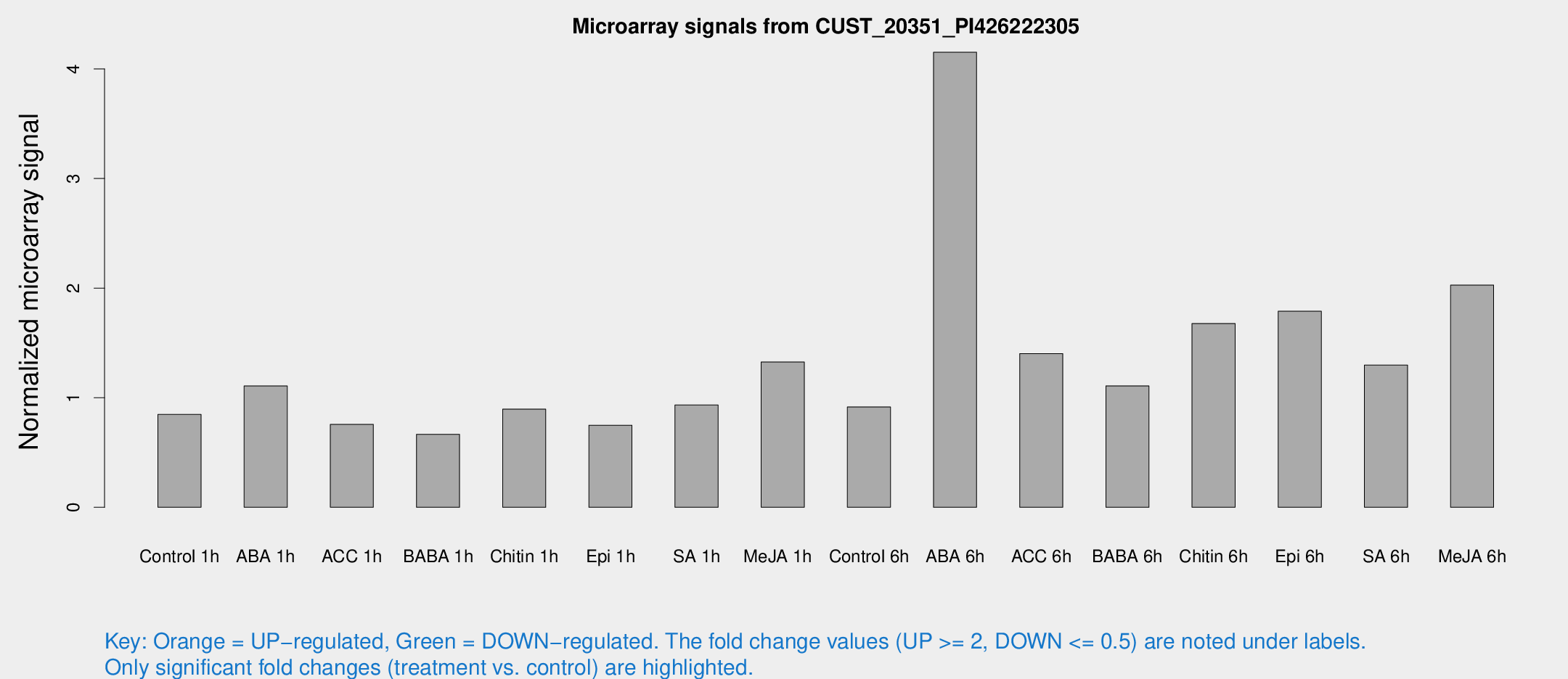
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 9.65352 | 3.59519 | 0.847275 | 0.378368 |
| ABA 1h | 10.2231 | 3.47753 | 1.10729 | 0.380335 |
| ACC 1h | 8.40403 | 3.96175 | 0.756432 | 0.382591 |
| BABA 1h | 6.66993 | 3.87648 | 0.665884 | 0.386305 |
| Chitin 1h | 9.23489 | 3.73566 | 0.895789 | 0.426728 |
| Epi 1h | 6.86271 | 3.57856 | 0.749273 | 0.400093 |
| SA 1h | 9.98606 | 3.65164 | 0.933719 | 0.342492 |
| Me-JA 1h | 12.2907 | 3.71717 | 1.32532 | 0.488837 |
| Control 6h | 9.85685 | 3.94521 | 0.916045 | 0.392943 |
| ABA 6h | 46.5683 | 6.05027 | 4.15225 | 0.439706 |
| ACC 6h | 16.9781 | 4.73472 | 1.40249 | 0.4198 |
| BABA 6h | 17.0547 | 9.32865 | 1.10781 | 0.643698 |
| Chitin 6h | 22.8785 | 9.30274 | 1.67725 | 0.907499 |
| Epi 6h | 21.262 | 4.60436 | 1.78881 | 0.403409 |
| SA 6h | 14.6339 | 4.17077 | 1.29757 | 0.616234 |
| Me-JA 6h | 22.3897 | 5.22055 | 2.0273 | 0.429359 |
Source Transcript PGSC0003DMT400049817 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G03090.2 | +2 | 2e-37 | 141 | 66/126 (52%) | methylcrotonyl-CoA carboxylase alpha chain, mitochondrial / 3-methylcrotonyl-CoA carboxylase 1 (MCCA) | chr1:739715-743819 FORWARD LENGTH=734 |
