Probe CUST_20048_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_20048_PI426222305 | JHI_St_60k_v1 | DMT400049195 | CATCTGTGAAACTTGGAACTCCGTGCTTATTTTCAATATAATGTTGGTATATCTCCATTG |
All Microarray Probes Designed to Gene DMG400019121
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_20048_PI426222305 | JHI_St_60k_v1 | DMT400049195 | CATCTGTGAAACTTGGAACTCCGTGCTTATTTTCAATATAATGTTGGTATATCTCCATTG |
| CUST_19967_PI426222305 | JHI_St_60k_v1 | DMT400049196 | TTGTGGCTTGTGGTGAAGGACTAAGGTCTTTGATGGCTATTTCAAAGTTACATGAAGGAG |
Microarray Signals from CUST_20048_PI426222305
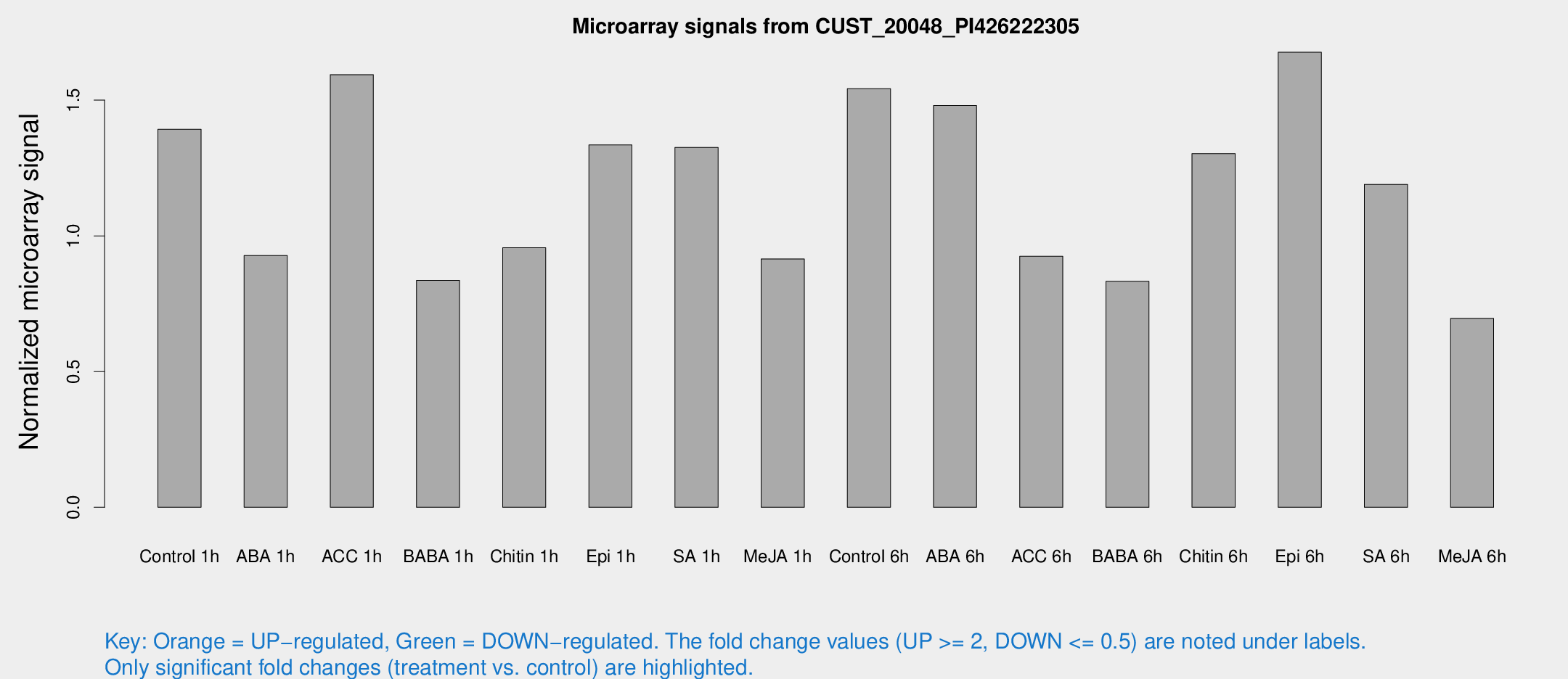
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 15.4051 | 5.77823 | 1.39244 | 0.536259 |
| ABA 1h | 8.27806 | 3.63229 | 0.927754 | 0.4567 |
| ACC 1h | 17.4667 | 5.81584 | 1.59403 | 0.545 |
| BABA 1h | 7.63349 | 3.94481 | 0.835973 | 0.437757 |
| Chitin 1h | 8.15302 | 3.78446 | 0.956145 | 0.451122 |
| Epi 1h | 11.9234 | 3.71592 | 1.33505 | 0.513126 |
| SA 1h | 13.8151 | 3.77698 | 1.32604 | 0.445628 |
| Me-JA 1h | 7.07186 | 3.78705 | 0.915038 | 0.493523 |
| Control 6h | 19.5344 | 10.6197 | 1.54249 | 0.891995 |
| ABA 6h | 15.7331 | 4.10486 | 1.48005 | 0.459865 |
| ACC 6h | 12.0862 | 5.52101 | 0.924816 | 0.446648 |
| BABA 6h | 9.07654 | 4.34233 | 0.832413 | 0.421706 |
| Chitin 6h | 13.9968 | 4.29418 | 1.30343 | 0.462119 |
| Epi 6h | 20.6373 | 7.53641 | 1.6766 | 0.845851 |
| SA 6h | 13.9655 | 6.98613 | 1.18979 | 0.570616 |
| Me-JA 6h | 6.52755 | 3.78591 | 0.695871 | 0.403295 |
Source Transcript PGSC0003DMT400049195 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G08920.1 | +1 | 1e-49 | 169 | 85/128 (66%) | Rhodanese/Cell cycle control phosphatase superfamily protein | chr3:2712274-2713117 FORWARD LENGTH=214 |
